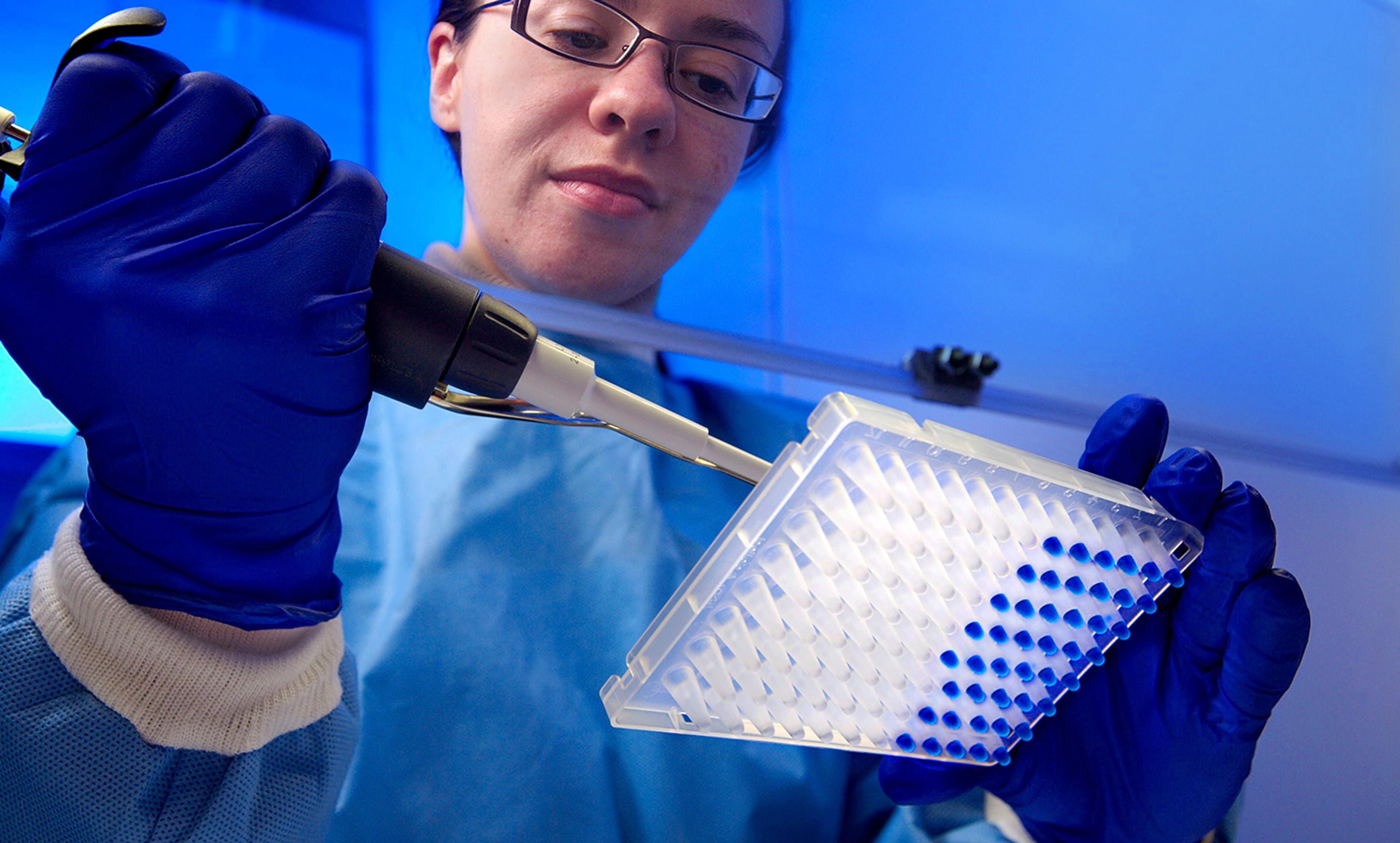
IAEA Imagebank/Flickr
The geneticist Kenneth Kidd started his career studying the family trees of purebred cows through their blood types A, B and O, each marked by a unique substance on the surface of red blood cells. By the 1980s, scientists had found genes coding for those blood types along with the specific variations in DNA that produce the distinct types. Scientists called the DNA that gave rise to such variant traits single nucleotide polymorphisms, or SNPs (pronounced snips). The insight was fundamental: a single nucleotide was the smallest DNA building block, and SNPs varied when one nucleotide was substituted for another. These single-nucleotide substitutions were the most common type of genetic variation among people. Before long Kidd had changed his interest from bovine genes to human ones. His groundbreaking work would ultimately allow humans to trace their ancestry around the globe and far back in time.
Now Kidd’s pioneering lab at Yale School of Medicine has enhanced the power of DNA analysis still further by identifying genetic markers with especially high resolution called ‘microhaplotypes’; these shorter strings of DNA are simply far less likely to degrade than the much longer strings used for analysis today, and stand at the front lines of forensics poised to help solve crimes. Aeon spoke with Kidd to learn how his new markers are helping to resolve two of forensics’ biggest headaches: degraded DNA and contaminated samples, where the DNA of multiple individuals is tangled together.
Why did you decide to look at short segments of DNA for forensic science?
I’ve been involved in forensics since the World Trade Center attacks [in 2001]. I was on the federal committee to advise about how to identify the victims’ remains. With only a segment of bone, you’ve got to use DNA, and it turned out that the standard forensic markers that were used for analysis then required longer segments of DNA. The current standard forensic markers are the short tandem repeat polymorphisms (STRPs) – segments of four nucleotides that repeat. [This is a more advanced technique than just comparing SNPs, often used in determining kinship.] The number of repeats differs on different chromosomes and in different people. We usually require at least 300 or 400 base pairs [to do an analysis]. But such long segments didn’t exist [at Ground Zero]; the DNA was often very degraded. It was said that the debris pile was at 2000 degrees [Fahrenheit] for weeks. DNA doesn’t like that. On the other hand, microhaplotypes are all 200 base pairs or less, so they have a much better chance of working on degraded DNA.
What kind of information can microhaplotypes provide today?
We can identify six or seven geographic regions of the world that are distinct. Imagine finding a skeleton in the woods: we have a reasonable probability with these of saying this is a person whose ancestry is South Asian or Native American or Far East Asian or European or African. That’s better and statistically more reliable than simply examining bones.
How do microhaplotypes compare to the current genetic markers, SNPs and STRPs?
They can do a better job than SNPs at determining family relationships. Say you have convicted criminals in a database, and you find DNA at a crime scene that is very similar to DNA in the database but it’s not identical: might it be a brother? A first cousin? The current forensic markers have high mutation rates, introducing uncertainty and confusion. As you look at more distant relationships, there’s a higher probability of a mutation, whereas the microhaplotypes have virtually no mutation rate. Mutations in the standard markers are as frequent as one in a thousand, whereas with the microhaplotypes it’s probably less than one in a million.
Microhaplotypes are much better at differentiating between two DNA samples at a crime scene. Why is that important?
If you touch something, you have left some skin cells behind, and if somebody else touches the same thing, some of their skin cells are there, so you’ve got DNA from two individuals now. In a lot of violent crimes there’s blood from the victims and there may be a little bit of blood from the assailant.
What forensic information will microhaplotypes ultimately reveal?
You can [use them to] learn about eye colour and skin colour and hair colour fairly readily. People are trying to find the genes responsible for the structure of the face to get a DNA photograph of a criminal, for example. They’re making progress but I’m not convinced it’s really good yet. Consider height: there are now well over 200 different genetic variants all over the genome affecting height. The most significant one results in only a few millimeters of difference. There are going to be a lot of genes that have to be considered to get any meaningful estimate of what the face is like. There are going to be genes that affect different bones, and then there’s the flesh on top of the bones. We don’t know why some people have big puffy lips and others have very thin lips, but that clearly affects how the face looks. Not to mention age effects. Ultimately, there will probably be some individual genes or groups of genes that will be useful, but it’s not going to be exactly photographic quality.
As more microhaplotypes are discovered and we get better at extracting information from DNA, do you envision any privacy issues arising?
I’m aware that a lot of people are very sensitive about this. I don’t think there’s a serious problem. The face is public, so is skin colour and eye colour. We’re not looking at anything that I think would be a privacy issue; we’re not looking at any causes of disease. There are disagreements about what is private. Is your ancestry private? In terms of forensics, catching a criminal, I have no problem seeing that the guilty person is convicted. I am extremely concerned about the innocent person being convicted, and that’s one of the reasons I’m doing this kind of research. All the convictions that are being overturned by DNA are a testament to the fact that the greatest value of DNA is in showing the innocent. The standard markers already serve that purpose, but we can easily exceed the current levels of accuracy with microhaplotypes.