In the mid-1980s, evidence started to emerge from labs across the world confirming that scientists were finally able to reach the nano level in experimental conditions and not just with their theories. Working at scales defined in millionths of a millimetre, Richard Smalley, Robert Curl and Harold Kroto reported the discovery of ‘buckminsterfullerene’ – a nanosized polyhedron, with 32 faces fused into a cage-like, soccer-ball structure, and with carbon atoms sitting in each of its 60 vertices.
These miniature ‘Bucky’ balls (named for their similarity to the geodesic dome structures made by the architect R Buckminster Fuller in the 1950s), are found in tiny quantities in soot, in interstellar space and in the atmospheres of carbon-rich red giant stars, but Kroto was able to recreate them in chemical reactions in the lab while visiting Rice University in Texas. Then, in 1991, Nadrian Seeman’s lab at New York University used 10 artificial strands of DNA to create the first human-made nanostructure, connecting up the DNA strands to resemble the edges of a cube, so marking the beginning of the field now known as ‘DNA nanotechnology’. Clever scientists with broad visions started to realise that a new kind of technology, prophesied by Richard Feynman in the 1950s, was finally materialising, as researchers achieved the capacity to visualise, fabricate and manipulate matter at the nanometre scale.
The term nanotechnology was coined in 1974 by the Japanese scientist Norio Taniguchi to describe semiconductor processes involving engineering at the nanoscale, but it entered public debate only with the publication of K Eric Drexler’s influential Engines of Creation (1986), a hyperbolic book of futuristic scientific imaginings of what might be achieved on the scale of the unimaginably small. Drexler’s book sparked longlasting controversies, notably focused on the weak scientific grounding of some of his ideas; but nothing stuck more to the public consciousness than his prediction of a hypothetical ‘grey goo’, scourge of a global dystopia involving out-of-control self-replicating machines devouring all life on Earth.
Perhaps the principal turning point in nanotechnology’s shift from theory to reality was the development of the scanning tunnelling microscope (STM), an ingenious machine that sidestepped the limitations of conventional light-based microscopes by interacting with nanostructures using a very sharp tip, as thin as a single atom. By scanning the atomic tip over the sample in a very controlled way, and mapping the local interactions of the tip with the exterior of the sample, an image might be created that delineated its surface with atomic precision. With the STM, which earned its inventors Gerd Binnig and Heinrich Rohrer the Nobel Prize in Physics in 1986, individual atoms became ‘visible’ at last or, more accurately, perceivable. But beyond imaging with unprecedented accuracy using a relatively simple, cheap tool, the STM had drastically new capacities: it was able to pick up and arrange atoms one by one.
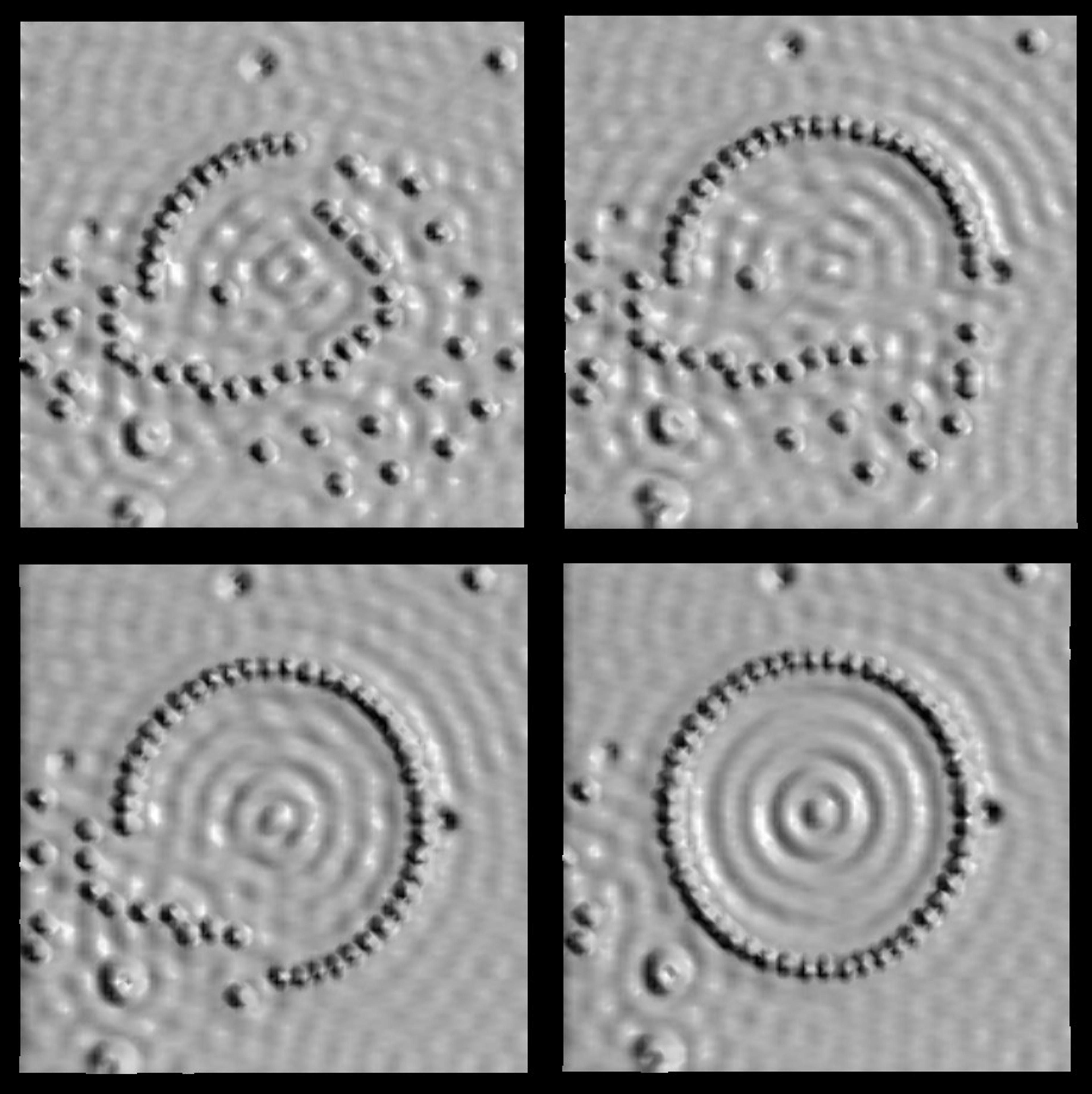

An STM image of iron atoms (the little spheres) on a copper surface (the smooth bumps are copper atoms arranged in a crystal). The STM tip is used to arrange the iron atoms in a circle, one by one. These images were taken by Don Eigler at the IBM Almaden Research Center in San Jose, California, in 1993. Image originally created by IBM Corporation
In the above image, we can see a standing wave appearing in the middle of the completed circle made with atoms. The ripple is produced by a wave of electrons emerging inside the quantum corral. This is a wonderful visualisation of quantum mechanics (particles behaving like waves) and it effectively summarises the state of the physics of matter at the time. As a reductionist tool, the STM could move and construct matter using single atoms as building blocks, and at the same time depict a complex wave phenomenon emerging as a result, which can be explained only by theories of matter that insist that ‘the whole is bigger than the parts’.
I saw this image when I was an undergraduate student reading physics in Madrid, and decided there and then that I had to find a way to be part of this world just starting to be explored. I am still at it but, like many of my contemporaries working in physics departments in the 1990s, I took my expertise and interest in physics at the nanoscale in a different direction: towards biology. From the start, nanotechnologists have been involved with and inspired by biology, primarily because the molecular players and the main drug targets in medicine – proteins, DNA and other biomolecules – are nanosized. But we physicists were also fascinated by the capacity of biology to produce materials that adapt, evolve, survive and even think – materials that surpass human technological abilities in every possible way.
Some of us were interested in why the Universe created life using elongated string-like molecules (polymers) able to fold in myriad nanoshapes (proteins) that assemble into all the living organisms that exist on Earth. The tools of nanotech enabled us for the first time to investigate, at the molecular level, the physical mechanisms that make life possible. Specifically, we could explore how the wonderful complexity of biology is underpinned by the dynamic behaviour of those biological nanomachines we call proteins. In the past 30 years, nanomicroscopes (mainly the atomic force microscope or AFM, a close relative of the STM) have revealed how proteins extract energy from the environment and perform the tasks needed to keep an organism alive. Far from being the static entities featured in traditional biochemistry books, proteins have been observed performing complex yet surprisingly familiar movements that sometimes bear an uncanny resemblance to macroscopic human-made machines. Some work as nanomotors that rotate to maximise the efficiency of chemical reactions; others can ‘walk’ on molecular tracks with a processive ‘hand-over-hand’ movement that allows them to transport cargos around the cell.
However, most attention was drawn to an area of research expected to bring quicker rewards: that of improving the effectiveness of medical treatments, especially in cancer chemotherapy. One of the main requirements for any drug to be effective is that it must reach its molecular targets in high enough concentrations. The problem with chemotherapy, though, is that it is very difficult to concentrate the drugs at the site of the tumour, hence the very high and toxic doses employed. In the 1980s, researchers found that nanoparticles accumulated at some tumours due to the peculiar structure of their blood vessels, which led to high hopes for using nanostructures to improve the delivery of drugs to cancerous cells. The field, instantly dubbed ‘nanomedicine’, grew rapidly. But expectations didn’t live up to the hype: currently fewer than 20 nanomedicines have been approved for use in cancer treatment. Seeking nano-powered magic bullets and lucky shortcuts to cure disease, while overlooking the complexity of the biology involved, has not proven especially fruitful. By mirroring the strategies of pharmacology, nanomedicine largely reproduced its failures.
Over the second half of the 20th century, the study of biological molecules and their chemistry turned into an increasingly structural pursuit, making biology itself a reductionist attempt to explain the whole by the sum of its parts. This trend was egged on by technological developments that led to the discovery of the molecular machinery of life: by mid-century, for example, scientists were able to resolve the detailed structures of biomolecules using X-rays on crystalline arrays of biomolecules extracted from their natural environment.
At the same time, progress in genetics and biochemistry led to a greater understanding of the chemical activity of proteins and their relation to the information stored in cellular DNA. Scientists were encouraged to develop an interpretation of biology that attempted simple solutions to its daunting convolutedness. They saw living organisms as biochemical computers executing a molecular program. And they viewed that program as an algorithm encoded in genes and materialised by proteins. Within this framework, medical researchers focused on identifying the rogue genes and proteins that caused diseases, and on finding drugs to deactivate them. The problem with this reductionist approach is that it doesn’t consider how biological cells, organs, tumours and organisms entangle themselves with their environment, combining and recombining, and collectively using their structures at every scale (from the nanometre to the metre and beyond) to keep on living, evolving and surviving.
To be fair, the reductionist approach to treating disease was justifiably fuelled by decades of revolutionary drug discoveries – antibiotics, chemotherapy and other ‘miracle drugs’ – that led to steep improvements in life expectancy. However, this century has seen a sharp decline in the number of effective new medicines produced. Between 2002 and 2014, a total of 71 new cancer drugs appeared, of which only 30 – found to prolong life in patients with solid tumours by an average of 2.1 months, compared with older drugs – have gained approval from the US Food and Drug Administration. The costly and largely ineffective trial-and-error methods used to identify new drugs, and the difficulty of conducting clinical trials, were partly responsible for this downward turn.
Undaunted by the dogmatism in molecular biology, they questioned reductionist models of life and disease
More significant, however, was the capacity of biological organisms to evolve resistance to new treatments. Antibiotic resistance is now widely identified as one of the biggest public health threats; meanwhile, the ability of cancer cells to build defences against chemotherapy has stalled pharmacologists searching for cancer cures. Bacteria and cancers are teaching us the same lesson that we are learning in other aspects of our relationship with nature: namely, that life resists reductionist approaches and bounces back with complex behaviours that thwart our optimistic strategies to dominate it.
Over the past two decades, both medicine and biology faced questions that the intellectual approaches and compartmentalisation of disciplines that prevailed throughout the 20th century could not resolve. In this sense, nanotechnology arrived at the right time, inspiring a new generation of physicists and engineers to turn their gaze towards biology. Pioneers generally look for new ways of doing things, escaping conservative establishments so as to build new realities, and so it was with the new generation of physicists and engineers. Undaunted by the dogmatism holding sway over molecular biology, they questioned reductionist models of life and disease. Their education gave them the tools of mathematics, the capacity to build machines to interrogate matter, and an eagerness to collaborate with materials scientists, enabling them to create new applications to transform the medical and technological landscape. Crucially, physicists carried the historical baggage of a field that, for the better part of a century, had been continuously forced to revise its core theories for interpreting reality. In physics, nothing was fixed for long any more. The success of general relativity and quantum mechanics revealed a reality far more wonderful and mysteriously complicated than we had dared to imagine. Who could say what might happen if biology were subjected to similar revolutions?
Consider for a moment how physics was transformed, from concentrating on how individual objects (atoms, planets) interacted with each other, to worrying about what happens when many objects interact with each other – atoms in solids, molecules in water – and pondering how seemingly unrelated patterns of behaviour emerge from those interactions. Macroscopic behaviour, such as superconducting or magnetic materials, was in this way explained by abstract concepts that extract the essence of how collective interactions between atoms give rise to simpler behaviour at larger scales (eg, magnets attracting each other). Similarly, soft matter physics has been able to predict the mechanical behaviour of large pieces of, say, rubber from the interactions of its molecular components (a development we now exploit in the design of high-tech running shoes and tyres).
Thanks to modern physics, we have finally started to discover how nature uses complexity to create the layers that compose our reality, from the Higgs boson to the behaviour of a flock of birds. The rules of the transformation from simple to complex, which then emerge in simpler behaviours again, are measurable, and can be modelled with mathematics. While biology is immensely more complicated than rubber (we are very far from explaining consciousness or emotion with maths!) nothing stands in the way of physicists trying to test the limits of our understanding of reality, and imagining what can be done with that knowledge.
One emerging field that exemplifies this transformation of science is ‘protein nanotechnology’, where nanotechnologists use proteins to design and construct microstructures and nanostructures, thereby imitating life. Proteins are the building blocks of life. In nature, they result from the careful and deterministic folding of molecular strings (polymers) consisting of combinations of 20 different units (amino acids). They can take on any imaginable shape and function at the nanoscale. In fact, we still don’t know how many different proteins are in our bodies (perhaps it is unknowable), since our cells could have the capacity to create and modify proteins as and when they are needed. Proteins work as light detectors in our eyes, electrical switches in our neurons, nanowalkers in our muscles, and rotary nanomotors to catalyse chemical reactions. They are responsible for detecting and reacting to the signals, forces and information from the environment in which an organism resides, and also for creating the structures that allow movement, the extraction of energy from food or the destruction of pathogens. No human-made artificial nanotechnology can dream of such capacities, but we can try to learn how life does it.
Protein nanotechnologists have been inspired by the recent success of scientists who have long tried to solve one of the thorniest conundrums of molecular biology: predicting the shape of a protein given information about the composition of its amino acid chain (this information is relatively easy to obtain). For years, the problem of prediction was considered too difficult, because the complexity of the interactions between the protein’s amino acids meant that their emergent shapes were impossible to calculate with the limited capacity of computers.
The first clear success in correctly predicting a protein structure came in 2014, with a team led by David Baker at the University of Washington. Two important factors can be singled out in spurring on this achievement. The first was crowdsourcing: Baker’s team successfully calculated the correct shape of the protein by exploiting the data-processing resources of thousands of volunteers and their personal computers. The second factor was realising that the interactions of the building blocks alone won’t do: correct protein structures could be inferred only by taking into account their evolutionary history. This was a radical departure from the way things had been done before. To understand the physics of biology, and then to construct structures as biology does, the evolutionary history of life on Earth must be included in the mathematical calculations.
This realisation came from the discovery made at Harvard by Chris Sander and Debora Marks of structural ‘staples’ (formed by amino acids sticking to each other) that hold a protein molecule together. Sanders and Marks looked at information contained in the genomic DNA of organisms that have proteins related to each other via a shared evolutionary history. When the staples are inserted into the computer model, it’s possible to explore how the protein folds within those constraints. The strategy, in other words, turns an impossible problem into a computable one. Unlike the inert structures of magnets, evolution guides complexity in biology, selecting the shapes and functions that allow survival.
Once scientists managed to predict protein structure, they immediately began using the ideas in reverse to create ‘designer proteins’ that don’t exist in nature and applying them to specific medical or technological purposes. To do this, they have to hack the molecular machinery of living microbial cells (mostly bacteria or yeast cells) and then re-engineer them ‘to produce’ the proteins they’ve designed on a computer. These ‘biological engineering’ technologies have made real one of the dreams of the nanotechnology pioneers: the deployment of molecular assemblers able to construct any shape with atomic precision, following a rational design.
They were inspired by how viruses kill bacteria by making nanoholes in their surface, using physics not chemistry
What’s interesting here is that the story has not unfolded quite as the pioneers of nanotechnology imagined, or as early visions of ‘nanorobots’ predicted, because this nanotechnology no longer emerges from a reductionist standpoint that envisages wholly artificial nanomachines deployed inside living cells. Instead, it utilises nature itself, harnessing its complexity and evolutionary history to create nanostructures. Inspired by medicine, this new approach to shaping matter is already producing astonishing breakthroughs, such as designer virus-like structures, with the potential to evolve and revolutionise the way we create vaccines or treat cancers.
This same approach is now being applied to the design of new antibiotics capable of overcoming bacterial resistance. Most antibiotics are small molecules that bind to bacterial molecules to kill them or prevent them growing, but bacteria can easily mutate to create chemical defences that rid them of the antibiotic. In December 2019, a UK collaboration between physicists, nanotechnologists, biophysicists, biologists, biomedical and computer scientists led by Maxim G Ryadnov of the National Physical Laboratory reported in the journal ACS Nano that they had constructed a nanoicosahedron using bits of proteins present in our immune system that could kill bacteria in a very efficient manner. The team was inspired by the way that viruses (like our own innate immune systems) kill bacteria by making nanoholes in their surface, using physics rather than chemistry. The nanoicosahedron they designed uses electrical charge and hydrophobicity to stick to and destroy bacteria, and it does this so fast that the bacteria cannot match its speed in evolving resistance to it.
The success of Ryadnov’s approach relies on a combination of skills: understanding protein structure, and the physics of protein assembly, being familiar with the biology of bacteria and viruses, and the biomedical techniques needed for assessing the effectiveness of antibiotics, with computational simulations and microscopy techniques from physics. It also points the way forward to a future where scientists can adopt evolutionary strategies (developed over time in our own immune systems) to overcome medical problems by engineering new versions of those strategies at the nanoscale.
Similar approaches have already enjoyed commercial success in fields situated far from medicine, such as electronics. A great example is the Californian biotech company Zymergen. With the aid of advance computing and AI, Zymergen engineers microbes (yeast and bacteria) to produce molecules that cannot be produced by traditional pharma or by chemical companies specialising in medical applications. But Zymergen’s reach goes beyond medicine: using a combination of biology, AI, computing and cutting-edge nanotech, the company has created the most advanced materials needed for the mobile phone industry. This February, Samsung unveiled a new-model mobile phone with a foldable screen. The main obstacle to creating such a device has been finding materials capable of matching the best in image quality, while simultaneously being pliable and amenable to folding and unfolding thousands, perhaps even millions of times during the phone’s life. Zymergen has shown that such high-tech materials are best constructed using computer-designed biological factories, fabricating at the nanoscale.
What the new nanotechnology seems to point toward is an inexorable dimming of the boundaries between the sciences. Though still in an embryonic state, the new transmaterial science of producing artificial materials inspired by biology is already being used to create new medicines, develop new strategies for regenerating tissues and organs, and improve the responses of the immune system. In parallel, hybrid bioinorganic devices that mimic biological processes will soon be used in new computers and electronic devices. By increasingly refining our ability to learn biology using the methods of physics, nanotechnologists are throwing off the yolk of reductionism, and learning how to distil the recipes of the Universe in order to fabricate and assemble matter from the nanometre scale up. In the process, they are revolutionising technology and medicine.