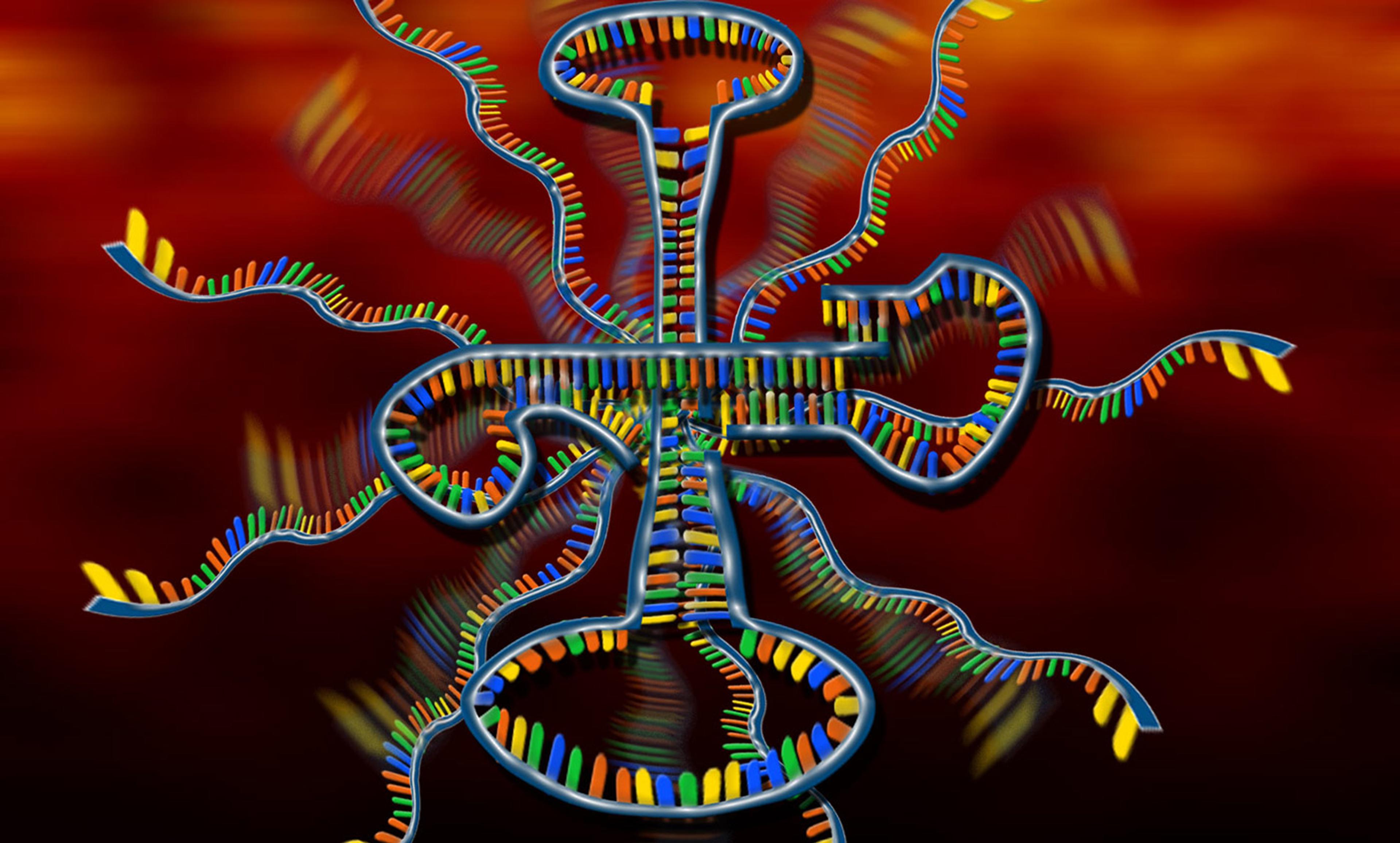
An artist’s rendering of a Ribonucleic Acid (RNA) molecule. Courtesy Nicolle Rager Fuller, National Science Foundation
What is life? For much of the 20th century, this question did not particularly concern biologists. Life is a term for poets, not scientists, argued the synthetic biologist Andrew Ellington in 2008, who began his career studying how life began. Despite Ellington’s reservations, the related fields of origins-of-life research and astrobiology have renewed focus on the meaning of life. To recognise the different form that life might have taken 4 billion years ago, or the shape it could take on other planets, researchers need to understand what, in essence, makes something alive.
Life, however, is a moving target, as philosophers have long observed. Aristotle distinguished ‘life’ as a concept from ‘the living’ – the collection of existing beings that make up our world, such as the neighbour’s dog, my cousin and the bacteria growing in your sink. To know life, we must study the living; but the living is always changing across time and space. In trying to define life, we must consider the life we know and the life we don’t know. As the origins-of-life researcher Pier Luigi Luisi at Roma Tre University puts it, there is life-as-it-is-now, life-as-it-could-be and life-as-it-once-was. These categories point to a dilemma that medieval mystical philosophers addressed. Life, they noticed, is always more than the living, making it, paradoxically, permanently inaccessible to the living. Because of this gap between actual life and potential life, many definitions of life focus on its capacity to change and evolve rather than trying to pin down fixed characteristics.
In the early 1990s while advising NASA on the possibilities of life on other planets, the biologist Gerald Joyce, now at the Salk Institute for Biological Studies in California, helped to come up with one of the most widely used definitions of life. It’s known as the chemical Darwinian definition: ‘Life is a self-sustained chemical system capable of undergoing Darwinian evolution.’ In 2009, after decades of work, Joyce’s group published a paper in which they described an RNA molecule that could catalyse its own synthesis reaction to make more copies of itself. This chemical system met Joyce’s definition of life. But nobody wanted to claim that it was alive. The problem was, it hadn’t done anything new or exciting yet. A New York Times article put it this way: ‘Someday their genome may surprise their creator with a word – a trick or a new move in the game of almost life – that he has not anticipated. “If it would happen, if it would do it for me, I would be happy,” Dr Joyce said, adding, “I won’t say it out loud, but it’s alive.”’
Joyce seeks to understand life by trying to generate simple living systems in the lab. In doing so, he and other synthetic biologists bring new kinds of life into being. Every attempt to synthesise novel life forms points to the fact that there are still more, perhaps infinite, possibilities for how life could be. Synthetic biologists could change the way life evolves, or its capacity to evolve at all. Their work raises new questions about a definition of life based on evolution. How to categorise life that is redesigned, the product of a break in the chain of evolutionary descent?
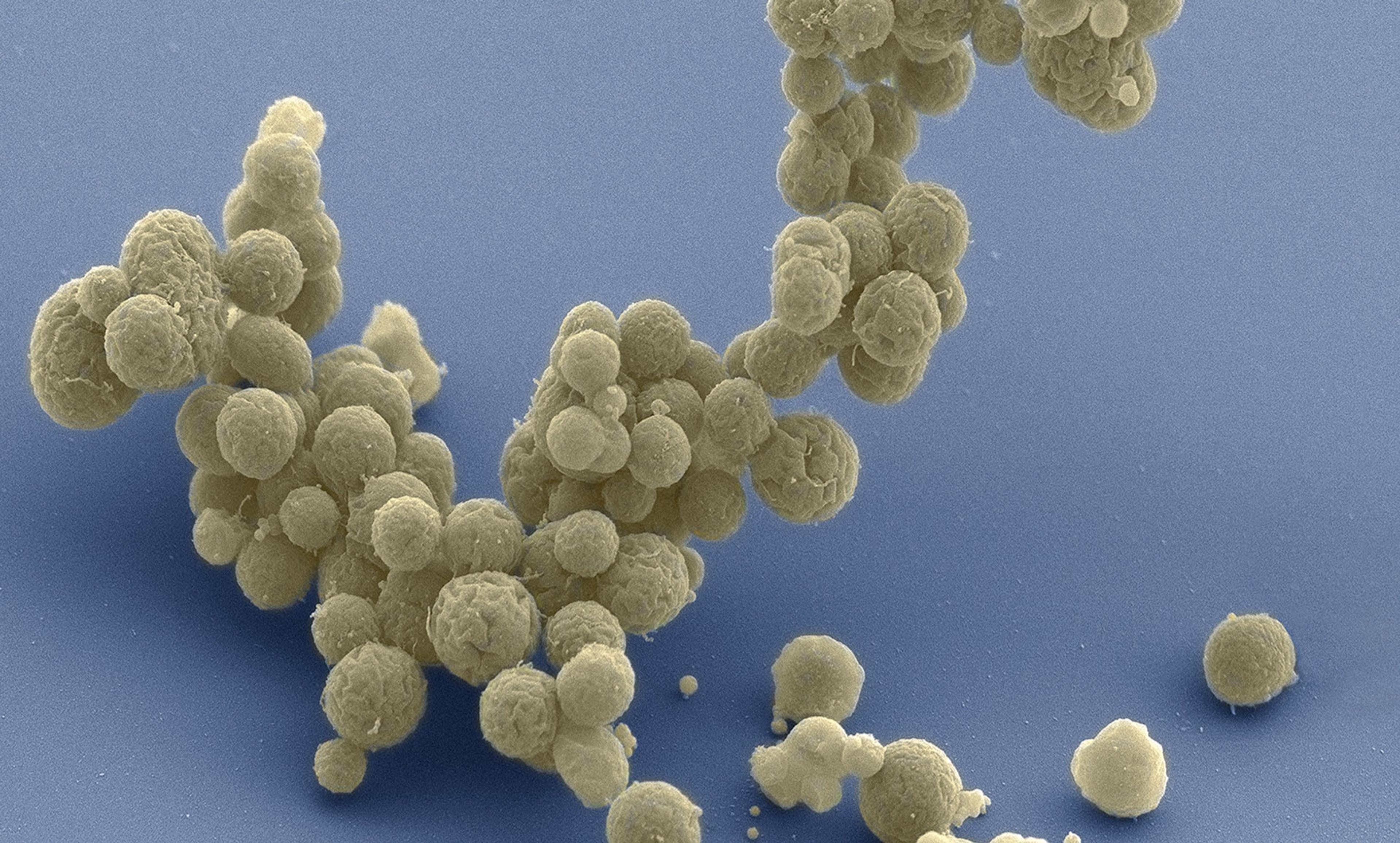
An origin story for synthetic biology goes like this: in 1997, Drew Endy, one of the founders of synthetic biology and now a professor of bioengineering at Stanford University in California, was trying to create a computational model of the simplest life form he could find: the bacteriophage T7, a virus that infects E coli bacteria. A crystalline head atop spindly legs, it looks like a landing capsule touching down on the Moon as it grabs onto its bacterial host. The bacteriophage is so simple that by some definitions it is not even alive. (Like all viruses, it depends on the molecular machinery of its host cell to replicate.) Bacteriophage T7 has only 56 genes, and Endy thought it might be possible to create a model that accounted for every part of the phage and how those parts worked together: a perfect representation that would predict how the phage would change if any one of its genes were moved or deleted.
Endy built a series of bacteriophage T7 mutants, systematically knocking out genes or scrambling their location in the tiny T7 genome. But the mutant phages conformed to the model only some of the time. A change that should have caused them to weaken would instead have their progeny bursting open E coli cells twice as fast as before. It wasn’t working. Eventually, Endy had a realisation: ‘If we want to model the natural world, we have to rewrite [the natural world] to be modellable.’ Instead of trying to make a better map, change the territory. Thus was born the field of synthetic biology. Borrowing techniques from software engineering, Endy began to ‘refactor’ bacteriophage T7’s genome. He made bacteriophage T7.1, a life form designed for ease of interpretation to the human mind.
Phage T7.1 is an example of what one synthetic biologist has called supra-Darwinian life: life that owes its existence to human design, rather than natural selection. Bioengineers such as Endy approach life in dualistic terms: a physical structure on the one hand, a pattern of information on the other. In theory, a perfect representation of life would enable a seamless transition between information and matter, intention and realisation: change some letters of DNA on your computer screen, print out an organism that looks and behaves just as you intended. With this approach, evolution threatens to corrupt the engineer’s blueprint. Preserving one’s biological designs might require making your engineered organisms unable to reproduce or evolve.
In contrast, Joyce’s desire for his molecules to surprise him suggests that the capacity for open-ended evolution – ‘inventiveness, pluripotentiality, open-endedness’ – is the critical criteria of life. In accordance with this idea, Joyce now defines life as ‘a genetic system that contains more bits [of information] than the number that were required to initiate its operation’. But according to this definition, given two identical systems with different histories – one designed and the other evolved – only the latter would be considered alive; the rationally designed system, no matter how complex, would be just a ‘technological artifact’.
Design and evolution are not always opposed. Many synthetic biology projects use a mix of rational design and directed evolution: they construct a host of mutant cells – variations on a theme – and select the ones that work the best. Although Joyce’s new understanding of life still involves evolution, it evokes the abrupt temporality of emergence rather than Darwin’s longue durée. Emergent life fits a culture of disruptive innovation whose ultimate ideal approximates something like the magic of pulling a kidney out of a 3D printer: the enchantment of joining together familiar things with new and surprising results. Design and evolution are also compatible when bioengineers look at genetic diversity as a treasure trove of design elements for future life forms.
For some synthetic biologists, the path to what the mystics called life-beyond-life – life that exceeds the living as we know it – now runs through biological engineering. Endy describes his vocation in terms of a desire to contribute to life by generating new kinds of ‘improbable patterns that continue to thrive and exist’. Joyce imagines life and technology joining forces against the fundamental thermodynamic tendency towards disorder and decay. What new forms life will take, only time will tell.