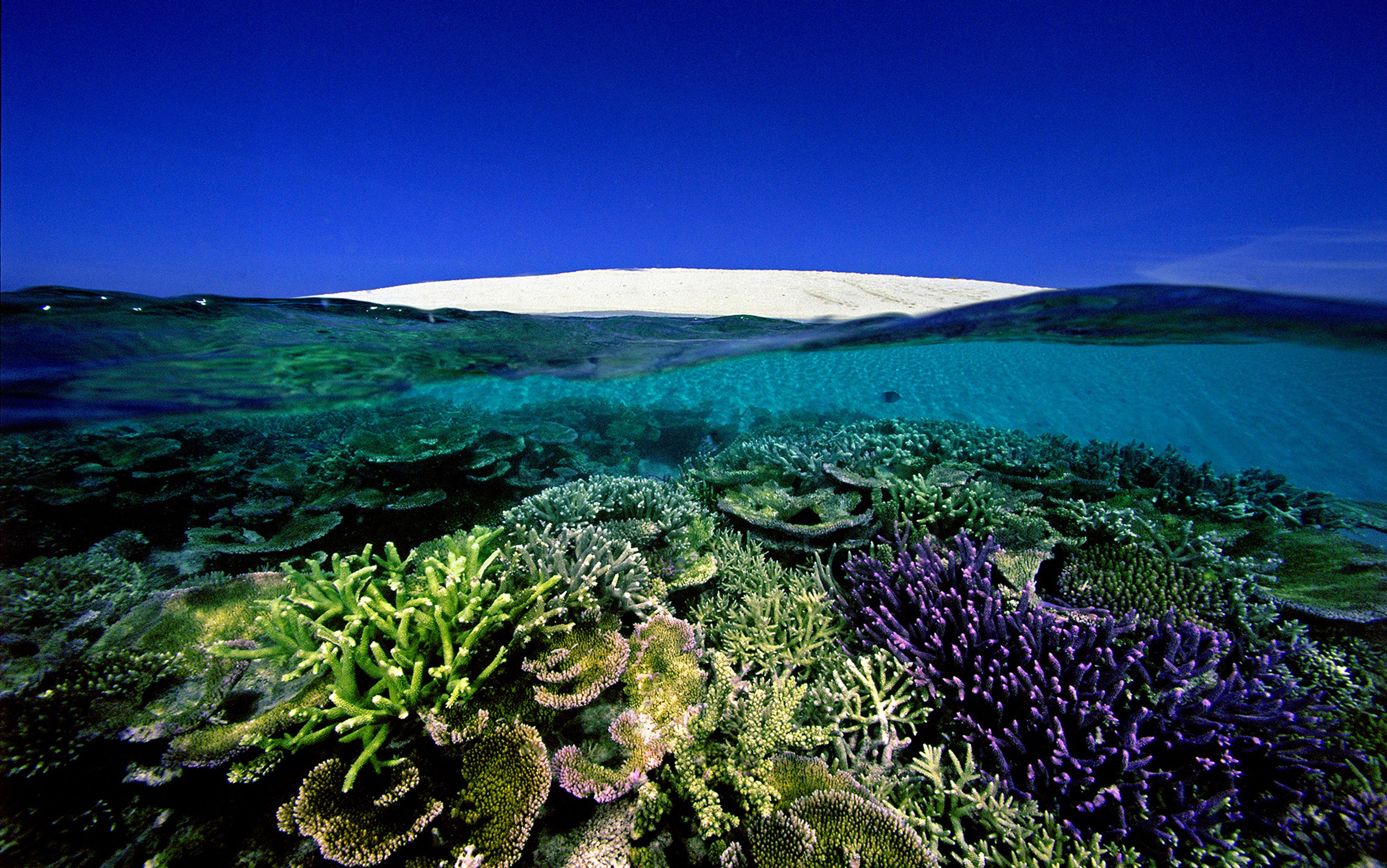
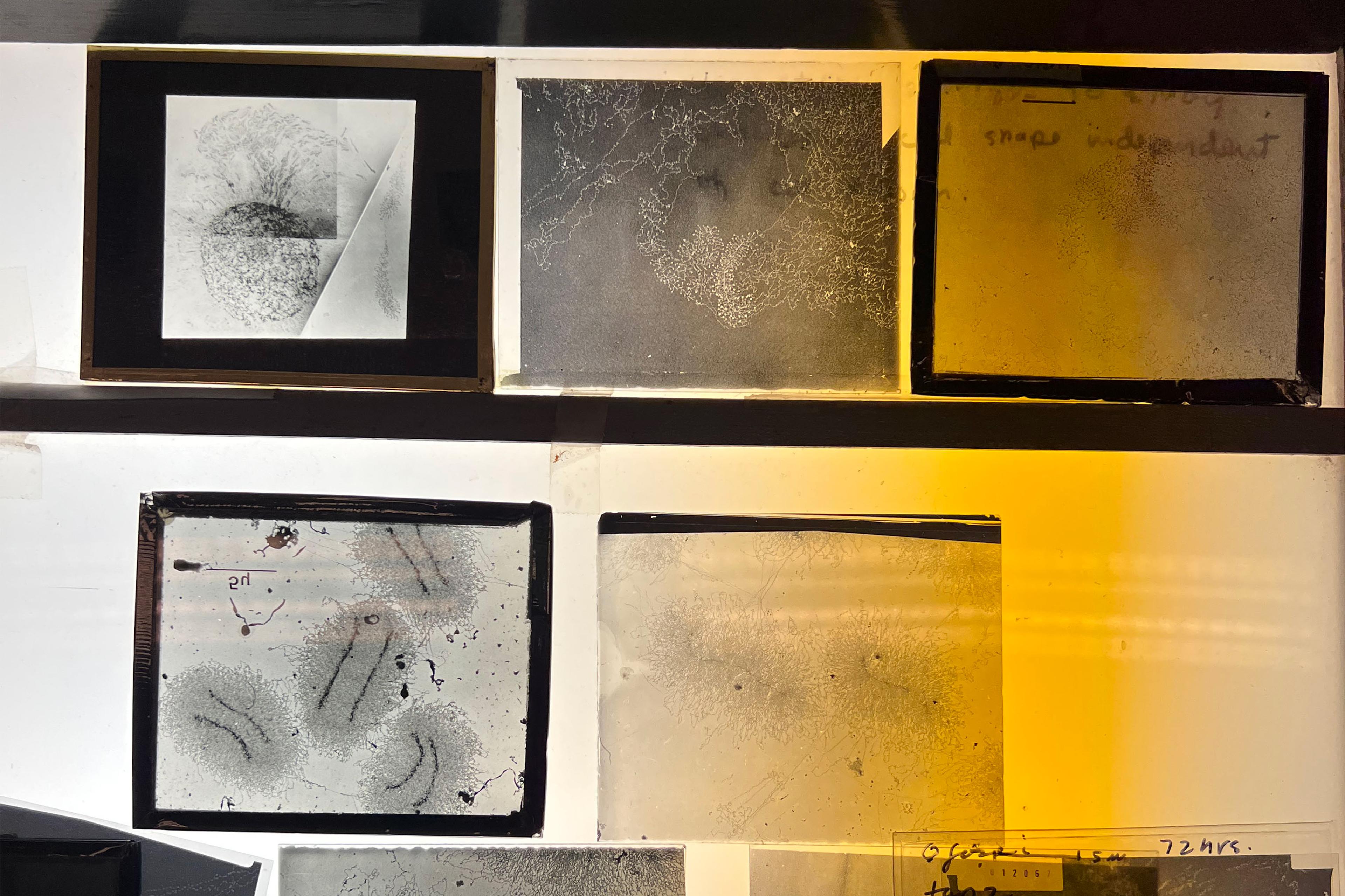
In the summer of 2017, two biologists set out for Denman Island in British Columbia, Canada, to build a house from scratch. This was not the first time. Years before, they’d built a cabin on the same 40-acre property, a cherished place of respite away from the lab. ‘We made the whole thing with almost no tools,’ Victoria Foe tells me with a playful, nearly mischievous energy. ‘We cut down the trees and made the joints.’ The only motorised device was a large drill the pair put in the refrigerator to cool down between driving dowel holes in the lumber. Foe, renowned for her precise studies of microscopic cells and embryos, debarked logs that would become the main framing posts. Garrett Odell, her life partner and close collaborator, designed the structure in a custom early CAD program he coded for Foe as a Christmas present.
But that summer things came to an abrupt halt. They had only just begun on the new house when Odell started feeling unwell. By October, he was diagnosed with aggressive liver cancer. ‘He died in May,’ Foe says. ‘It made the light go out in the world.’
For Foe, a developmental biologist, life and biology are intimately entwined, each motivating and deepening understanding of the other. Rather than turn her away from science, Foe’s grief pulled her back to an observation about the fragility of life she’d carried for decades, ever since the start of her PhD in the late 1960s.
Then a graduate student at the University of Texas at Austin, Foe was studying the shape of genes in milkweed bug embryos no larger than a grain of rice. Foe took embryos and burst their cells, releasing deoxyribonucleic acid (DNA) from nuclei, the inner compartments where genes are stored. After centrifuging this liquified material onto a microscopic grid, she coated it with platinum, forming a thin, heavy dusting of metal that would make genes show up under an electron microscope. In the resulting micrographs, she saw genes that were shockingly long, stretching to nearly 20,000 ‘letters’ of the genetic code built from four chemical bases labelled A, T, C, and G.
By the time Foe got her first glimpse of genes, the meteoric rise of molecular biology was well underway. This momentum was fuelled by evidence for the ‘central dogma’, which states that DNA stores the ‘code’ of life; that this code is transcribed into ribonucleic acid (RNA); and that RNA, in turn, translates the code into the chains of amino acids forming proteins.
Placing her own vulnerability within that of the living world, she was struck by the fragility of genetic material
First proposed by Francis Crick in the late 1950s, the central dogma of molecular biology was popularised in James Watson’s textbook Molecular Biology of the Gene (1965); there, Watson added that the information transferred from DNA to protein can flow only in one direction. The idea that DNA’s value lay solely in the proteins it coded for led, ultimately, to the realisation that the vast majority of DNA in our cells didn’t code for protein at all; this ‘non-coding’ DNA was thus considered dark matter, a nuisance tolerated by complex organic life for more than 2 billion years – probably since a retrovirus invasion lodged this extra material into our single-celled ancestors’ genomes. Biologists said it was junk.
But when Foe observed the enormously long threads of DNA back in Texas, she sensed there might be more to the story – but what? Placing her own vulnerability within that of the living world, she recalls being struck by the fragility of genetic material, two helical threads of nucleic acid that can be severed clean through every time the cell containing them divides. The more thread, the more opportunity for breakage, which is a cause of many cancers. In multicellular organisms such as humans, the genes inside our cells are much longer than they seemingly need to be. Of the 10,000 loops of DNA packed into 23 pairs of chromosomes (tight, spool-like bundles of genetic material) within nearly every human cell, only about 1 per cent is used to make proteins. If the expanse of our DNA exposes it to damage, Foe wondered, then why have we evolved this way? Far from ‘junk’, she proposes, the fragile stretches of non-coding DNA in our cells may enable our very existence.
Foe is best known for work she did in the 1980s at Friday Harbor Laboratories, University of Washington, charting where and when cells divide in the earliest stages of the fruit fly embryo – an organism that has anchored modern genetics. During the first two hours of the embryo’s life, its nuclei divide in lockstep. Soon, that unity breaks: distinct rhythms of division set in, with groups of cells falling into their own tempos as the embryo begins laying the foundation for different parts of the body to develop. The timing and locations of these dividing groups, Foe realised, were highly predictable. She detected this through meticulous observation of hundreds of micrographs, in which she stained every cell so its inner scaffolding was visible, and mapped these division domains in full-page spreads of colourful drawings and micrographs.
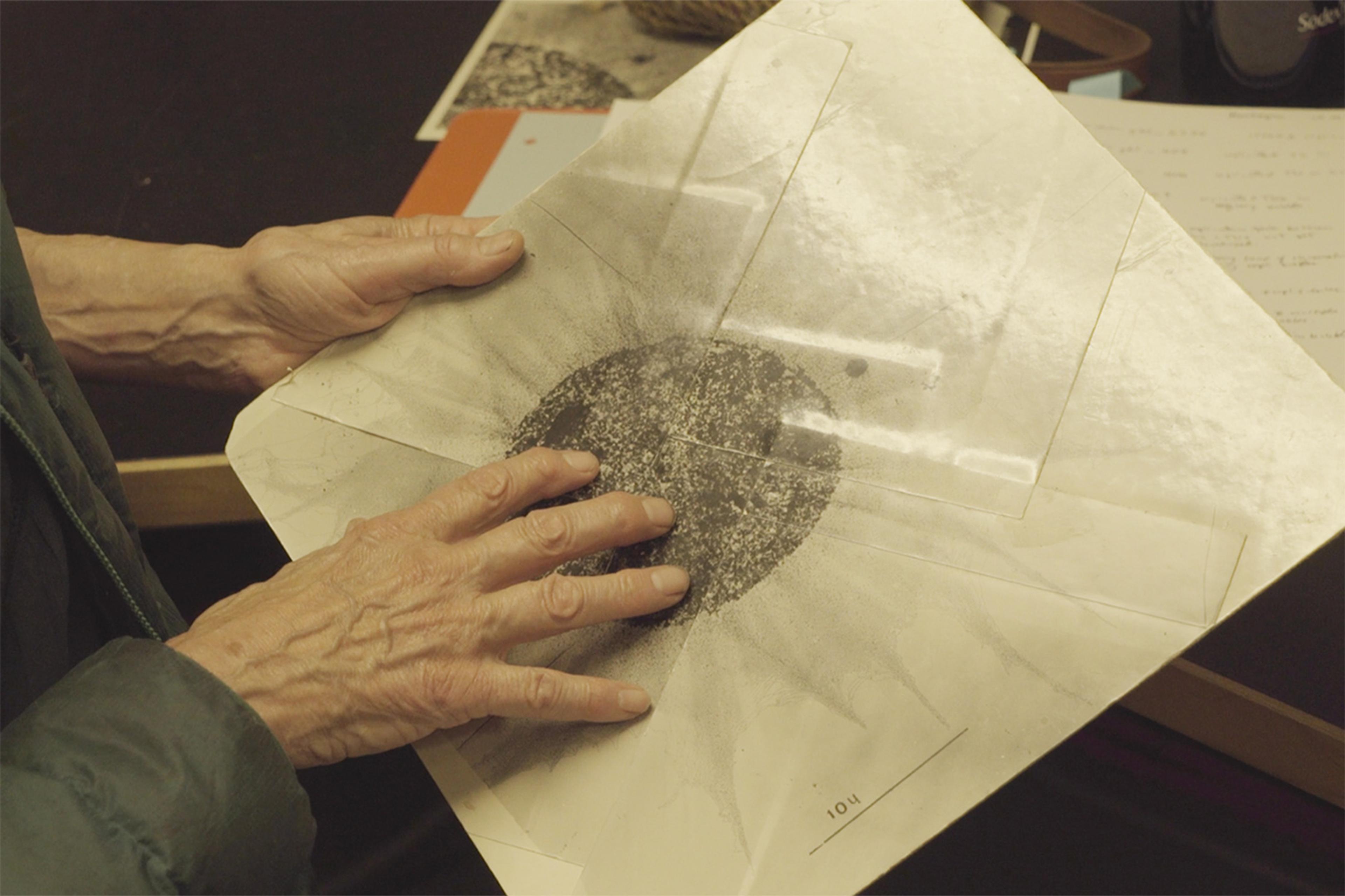

Victoria Foe holding an electron micrograph collage of a nucleus with DNA spread at its edges. Photo by Beatrice Steinert, 2023
To work on this project uninterrupted by the usual demands of academic science – teaching, mentoring trainees, and managing labs with often enormous overhead – Foe, who did not yet have a faculty position, convinced the US National Institutes of Health to give her a personal grant. She has held fast to this model of being a biologist ever since. This commitment made space for an attention to her craft that has earned her Guggenheim and MacArthur ‘genius’ fellowships, among other accolades. Most importantly for Foe, it has kept her close to what she holds dear: ‘Making something beautiful is really, really important to me.’
On a rainy early fall day, I meet Foe in her lab on the coast of Friday Harbor on San Juan Island, Washington State. For someone of her stature, she is surprisingly nervous, which I quickly realise emanates from her desire to be certain I completely understand the story she has to tell. She is wearing hiking boots and a light down jacket, attire equally suited for long days in the lab and walks with her dogs along the rocky northern shores of San Juan. Following Foe’s quick movement from the front room with a view of the San Juan Channel to the windowless back room is arresting. The walls of the space are covered with large prints of her micrographs and drawings, reminiscent of the elaborately illustrated wall charts that adorned 19th-century scientific lecture halls. Rather than the digital screens that fill most contemporary labs, negatives are spread out on lightboxes. The centrepiece of the room is Foe’s towering electron microscope that she refers to with the endearment of an old friend.
‘This is a nice one,’ she says with tenderness, ‘it’s laid out nice and relaxed, not pulled or under tension’
Foe’s interest in embryonic cellular rhythms originated in her graduate work imaging the architecture of genes, a practice she has returned to in the past decade, now using fruit flies as her model organism. After chemically fixing fruit fly embryos – freezing them in time – she spreads open their nuclei and coats the resulting tangle of genetic material with platinum. She then loads the preparation onto the electron microscope and begins looking for patterns. She can spend up to a month observing a single embryo. Kept company by audiobooks, Foe spends hours on end detecting the looping and branching forms that appear when genes are actively being read to produce RNA during development. When she finds something interesting, she begins imaging it piece by piece to collage together an expanded field of view.
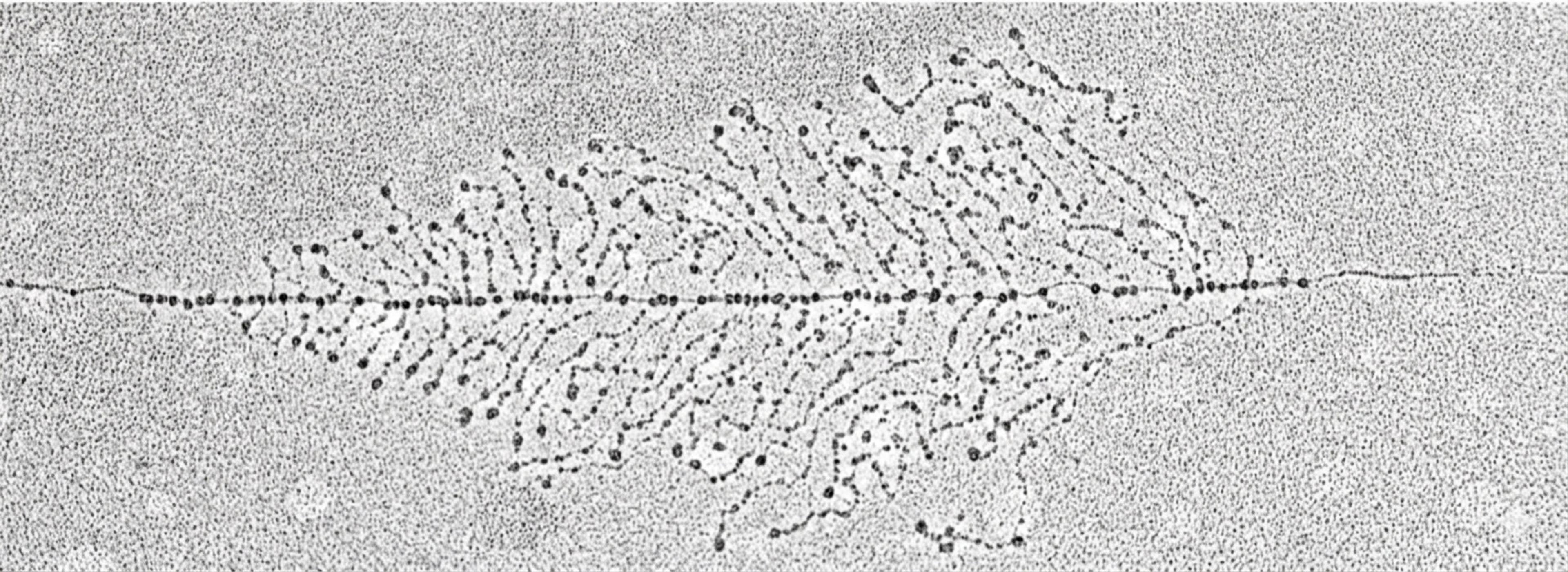
Electron micrograph of a fruit fly (Drosophila) gene. Courtesy of Victoria Foe
At the microscope, Foe pulls up one of her micrographs to show me. ‘This is a gene,’ she says, gesturing to a five-micrometre length of thread. The gene she encompasses with outstretched fingers has been dislodged from the nuclear tangle just enough to reveal its shape. ‘This is a nice one,’ she says with tenderness, ‘it’s laid out nice and relaxed, not pulled or under tension.’
Delicate branches extend perpendicularly from the base of the gene. These are nascent fibres of messenger RNA, built base by base from the underlying DNA template. Higher up, the branches lengthen, reaching out with knotty twists and turns. Foe points to the specialised proteins – called spliceosomes – encrusting these branches that cut away non-coding stretches from the newly transcribed RNA and stitch the remaining pieces together. The result is a cleaned-up message, an RNA transcript ready to leave the nucleus and travel into the rest of the cell, where it can be used to make proteins. These mature transcripts, Foe says, ‘get to the end, fall off, and off they go.’
As a graduate student, Foe hand-traced images like these to unspool them into meaningful parts, laying a foundation for our current understanding of how genes function. What she found was that there were distinct locations where DNA-to-RNA copying began and ended, bounding what she called a ‘transcription unit’ – the entire stretch of DNA that was transcribed – or rethreaded – to activate it. Foe noticed in her drawings, published in Cell in 1976, that most of the transcription units had the same undulating architecture: small branches near the start site, followed by long ones, and ending with RNAs that were much shorter than the full length of DNA from which they were made.
Milkweed bugs, it turned out, were no exception. Over the decades, biologists found long transcription units – more commonly called genes – in species after species. Many of these genes are studded with introns: stretches of DNA that are copied into RNA only to be cut out before proteins are made from them. In humans, this genetic extravagance is taken to an extreme. The largest human transcription unit spans more than 2 million base pairs and would take at least 17 hours to transcribe.
As these discoveries were being made, biologists considered the system grossly inefficient. For many of Foe’s colleagues, she recalls, ‘that this non-coding stuff was being read into RNA seemed metabolically incredibly wasteful’. Together with other non-protein-coding stretches, the introns that make genes such as our own so long were called ‘junk DNA’ because they didn’t appear to contribute anything meaningful to an organism’s biology.
‘But it makes a lot more sense when you can understand that it’s a timing mechanism,’ Foe tells me. Her latest paper, published in 2022, distils a lifetime of work synthesising how gene activation and cell division are integrated to create wings for flying or finely branching roots for absorbing nutrients from the earth. Foe argues that the degree of complexity in a multicellular species depends in part on the choreography made possible by the length of their genes, including their introns. Embedded in the rhythm of cell division, these extra stretches of transcription add a layer of time – one that helps make room for elaborate body plans and dynamic nervous systems.
Inheritance can look beautifully straightforward – until you ask how genes actually contribute to making organisms. Gregor Mendel – the 19th-century Austrian monk who helped inaugurate what the historian of science Evelyn Fox Keller called ‘the century of the gene’ – caught nature at its most cooperative. When he crossed pea plants of different heights, the outcomes were a crisp, teachable ratio: in the next generation, roughly three-quarters of the offspring were tall and the remaining quarter short. Plant height, Mendel reasoned, had to come from a discrete factor – something passed from one generation to the next and shuffled according to simple combinatorial rules.
That tidy calculation offered a dream of biological clarity. The simplicity of Mendel’s scheme allowed him – and then a century of 20th-century biologists – to begin naming heritable factors that, to a certain degree, set the range of forms an organism can become. But the deeper researchers looked, the more Mendel’s clean, straight-line causation began to fray. In complex life, the same genes can lead to wildly different outcomes – not because the DNA changes, but because the use of DNA within cells changes.
In 1961, the molecular biologist Jacques Monod proclaimed that ‘anything found to be true of E coli must also be true of Elephants.’ While this holds true for the basic DNA-to-RNA-to-protein logic of genetics, it loses meaning in the context of development. In eukaryotes – which include multicellular plants, animals, and fungi – traits seldom emerge from a lone gene acting like a switch, as operates in the bacteria E coli.
As an organism of one cell, E coli, a prokaryote, has only a single copy of its genome propagating signals throughout its body. An adult elephant, in contrast, has more than a quadrillion cells, nearly all of which house its own set of every gene in the genome. It is not just that genes, or specific sequences of DNA, are strung together (a staggering number are shared across the tree of life) but – even more potently – when, where and to what degree they are set in motion in cells that makes the difference. Starting from a single fertilised cell, the single-celled embryo, cell divisions gradually expand and sculpt multicellular bodies. As development flows from one moment to the next, cells take on new behaviours and identities that are initiated by interacting collaborations of genes and cellular processes: bone cells begin secreting minerals to counter gravitational forces, neurons make connections at their synaptic junctions, and heart cells electrically pulse in unison to disperse oxygen. An ultraprecise mechanism is needed, then, to bring on each gene in the right place and at the right time in a constantly transforming multicellular landscape.
In the 1950s, the organism was viewed as a one-dimensional machine to be decoded
The length of genes, their physical presence in the cell, Foe proposes, acts like a molecular clock – a built-in timer that tunes exactly when, whether and how much RNA – and ultimately protein – is made during a given cell division. The moment of division is key. During development, cell division is not simply growth; as cells divide, they also frequently change identity, turning different genes on and off so that one cell becomes muscle, another nerve, another bone. Those identity shifts depend on bursts of RNA production that precede the making of proteins.
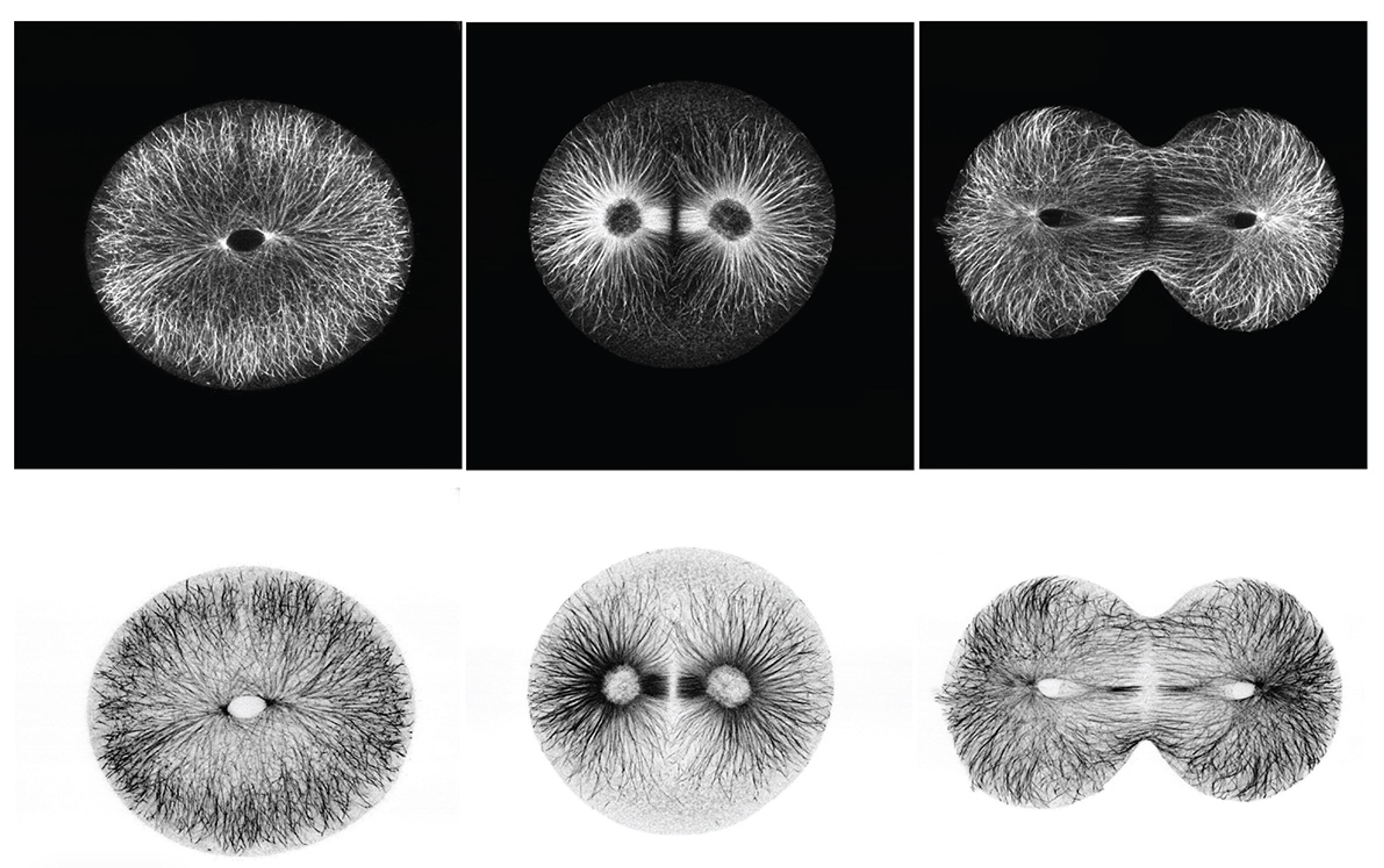
Strongylocentrotus droebachiensis (sea urchin) cell-division series, 2008. Courtesy Victoria Foe
The reason gene length can function as a timer is built into the ins and outs of cell division. Every time a cell divides, it has to package its DNA for safe distribution. To do that, it condenses the DNA within its nucleus into chromosomes that can be moved and evenly divided between daughter cells. As the genetic material is packed for division, RNA polymerases – enzymes that move along genes, reading the code and building an RNA copy – are knocked off the DNA. RNA construction thus grinds to a halt. The timing of this intricate process, known as the cell cycle, is itself variable and carefully controlled.
In fruit flies, for example, the hormone that initiates metamorphosis floods the larval body in a punctuated pulse. Multiple genes involved in this dramatic transformation from ground-crawling to flying life await their cues. But even after the signals come, beginning the process of transcription, it takes time to assemble the required RNA nucleotide by nucleotide. If a short gene takes five minutes to be copied, its RNA will appear on the scene swiftly and in copious amounts during a two-hour cell cycle. A longer gene unit full of DNA that needs to be cut out, however, will be finished only by the very end. If a gene is extra-long, its transcription will be halted midway; its RNA message will be finished for protein translation only in cells with a longer cycle. Once the cell has split in two, the clock resets.
This finding exposes the limits of a century of biological thought – not just the simplicity of Mendelian genetics, but also a Cold War impulse to recast genes as circuitry. As the historian of science Lily Kay has documented, beginning in the 1950s, computational metaphors became the bread and butter of molecular biology; the organism – at first, mostly viruses and single-celled bacteria – was viewed as a one-dimensional machine to be decoded.
We are not merely machines governed by switches, Foe suggests, but dynamic matter
By the 1970s, developmental biologists had expanded this framework into multicellular beings, elaborating the linear diagrams of Monod and his colleagues as they experimentally parsed gene regulatory networks (GRNs), cohorts of genes with the ability to turn one another on or off among groups of cells. Abstractions of a kind of information flow, these diagrams eclipse the physicality of the cellular milieu in which genes are made meaningful.
The informational gene has held enormous sway in the life sciences over the past half-century. DNA as ‘code’ – as if life could be read from a single text – proved a profoundly seductive idea, one powerful enough to reshape institutions as well as research agendas. In the mid-20th century, biology departments split along these lines, as molecular biologists, led by figures such as James Watson, pursued the logic of genes as information, while other biologists remained focused on organisms, development and evolution.
Explanations centred on ‘master’ genes became especially compelling. Technologies built around isolating, sequencing, synthesising and editing DNA transformed what counted as experimental progress, making it straightforward to remove or amplify a gene and document the discrete changes resulting from that perturbation. In fact, this strategy has been extraordinarily productive; the volume and speed at which genetic data is published continue to expand at a breathtaking pace. Even the recognition that genes operate within complex regulatory networks did little to dislodge the deeper assumption that biological meaning resides primarily in information coded in DNA sequences.
Foe’s work presses against that assumption. Genes are not only messages but material entities embedded in time, subject to interruption, constraint and decay. Living systems, especially those of multicellular organisms, cannot be understood solely as circuitry or code. Instead, development unfolds through the physical realities of our molecular building blocks, colliding, being halted, and beginning again. We are not merely machines governed by switches, Foe suggests, but dynamic matter – sensitive to our surroundings, vulnerable to damage, and open to transformation.
When I first learned about Foe as an early graduate student, I was still immersed in the distancing ethos of most of biology. Against this backdrop, I was instantly captivated by her sense of deep interiority, and her work grabbed me with a force I had never experienced from scientific literature. In an age when scientists strive for efficiency, Foe’s papers are long and lusciously dense. They are also by contemporary standards unusually visual, with clear and detailed images often laid out in full-page spreads. Her presence jumps out in every turn of phrase, particularly in her methods sections, which elaborate her mastery in wrangling the smallest of living beings under the microscope.
Foe’s trajectory has been unique, especially as a woman who began her scientific training in the 1960s and is a lifelong women’s rights and antiwar activist. ‘I did not have in my mind that I wanted to be a scientist,’ Foe says when I ask about her early years. But the British education she received living abroad as a teenager seeded her interest in biology when she began as an undergraduate at the University of Texas at Austin. Supported by a US mandate to invest in science during the Cold War, she received a prestigious fellowship for graduate school. She began a zoology PhD in 1968 but immediately had trouble finding an advisor. At universities across the country, Foe recalls, ‘the assumption was it’s just not worth training women because they’re going to get married and they’re going to get pregnant, in some order.’ It was obvious to her that access to birth control was essential for increasing equity in science. Along with her lab-mate Judy Smith, one of the only other women in her department, and another student, Barbara Hines, Foe started a birth control information centre near campus and went on to be involved in the state legislative work that would give rise to Roe v Wade.
‘Once you get into [it], there’s just one more delicious puzzle after another’
It was during this time of upheaval across the US that conflict began to emerge in university biology departments, including Foe’s, over whether or not DNA was the prime mover of life. Departmental infighting is what caused her PhD advisor to move to the University of Washington. Foe eventually followed him there so she could finish her degree, driving from Austin to Seattle in a rickety purple VW Beetle she borrowed from an ex-boyfriend. After a stint in San Francisco, where she first began working on fruit fly development at the University of California, Foe returned to the Pacific Northwest in the 1980s. She took up residence at Friday Harbor Laboratories, a nature preserve and marine research station on San Juan Island, where she has remained ever since, adding studies of myriad marine embryos to her zoological repertoire.
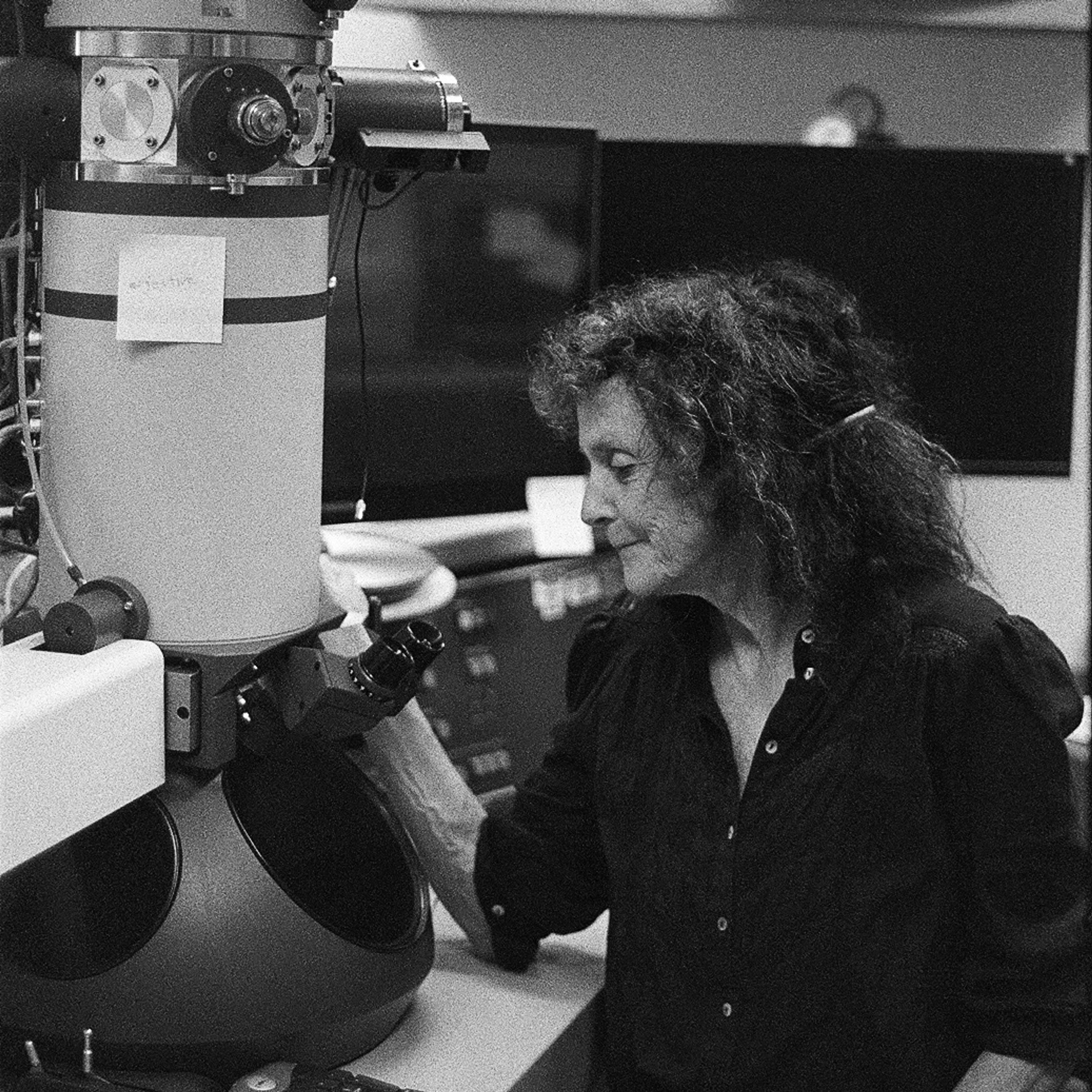

Victoria Foe at her electron microscope. Photo by Beatrice Steinert, 2023
In several respects, Foe’s work and career resemble that of an artist even more than a scientist. On her research webpage she writes: ‘My favourite research instruments are my eyes.’ Her work is forwardly descriptive and relies on the same observational and spatial skills of visual artists. While descriptive morphological studies are regarded by many biologists as a lesser way of knowing – well below the rigour of molecular experiments – they are in fact, as the historian of biology Soraya de Chadarevian has argued, the foundation of modern life science. Also like artists, Foe embraces her subjectivity. Much of modern science is about conformity: scientists strive to cultivate the look and sound of removed authority, especially in their publications, because that is how they maintain credibility. In The Politics of Women’s Biology (1990), Ruth Hubbard – the first woman to be tenured in Harvard’s biology department – writes that ‘scientific writing implicitly denies the relevance of time, place, social context, authorship, and personal responsibility’. Foe has largely disregarded these norms, putting herself – her style, her ‘I’ in her writing, and her valuing of visual aesthetics – unapologetically into her work.
When I posit Foe’s work as art to her, she seems somewhat uncomfortable with the proposition, even while her image-adorned lab around us suggests otherwise. I sense this stems from the wedge driven in the past century between art and science; also, at least during Foe’s formative professional years, a whiff of her artistic inclinations would have detracted from her being seen as a serious scientist. Being a woman scientist, or a woman artist, itself was hard. A woman artist-scientist? Too many boundaries crossed. I also gather, however, that Foe doesn’t think much about how her work is categorised by others. Her primary motivation comes from within. ‘I got into science because an opportunity was made available to go to school. And then once you get into [it], there’s just one more delicious puzzle after another … It’s just so intoxicating understanding how the world works, what life is, you know. Who wouldn’t want to study that?’
When I first searched for Foe online, I found an image of her and Odell in an embrace. They are crouched on the ground, surrounded by sun-raked trees. Their arms are slipped into one another’s sleeves, and their similarly textured hair entwines as Foe’s cheek rests on Odell’s forehead. This photograph accompanies the Friends of Foe webpage that raised funds for Foe to finish the research she stepped away from while caring for Odell when he got sick. It poignantly captures the price of complexity, of winding entanglement with the world. Present in this snapshot is the possibility that life brings – of beauty, love, companionship – but also the chance that it can unravel.
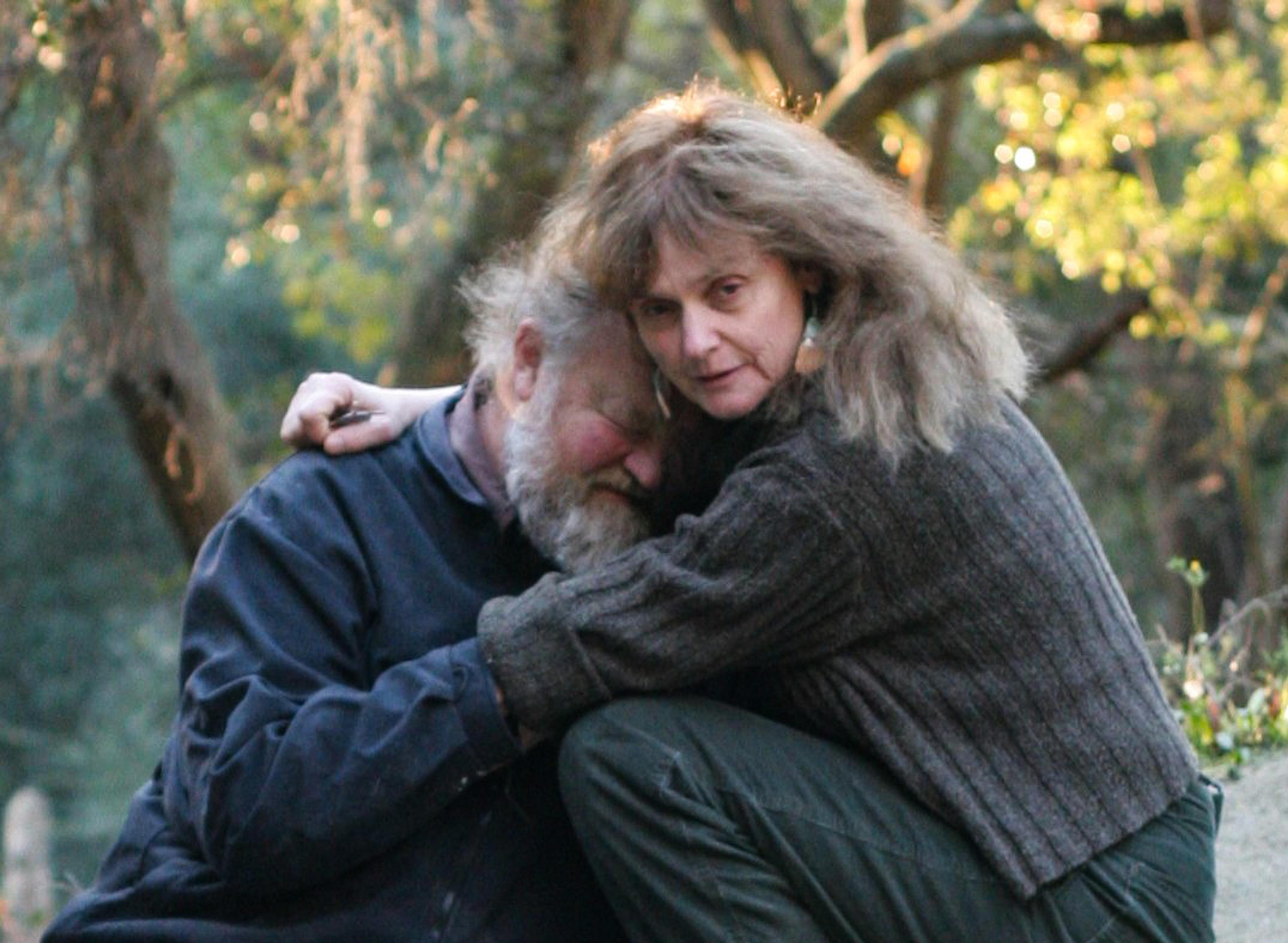

Garrett Odell and Victoria Foe. Courtesy of Friends of Foe Fund, University of Washington
Foe and Odell first met at Friday Harbor in the late 1980s, soon after Odell was recruited by the University of Washington zoology department. Trained as a mathematician who began his career at the Rensselaer Polytechnic Institute in New York, Odell’s fascination with how molecules create organisms drew him into biology. His early work on mathematical models of emergent behaviours – ranging from insect predator-prey interactions to cytoplasmic streaming in slime moulds – anticipated the rise of systems biology and the computational simulation of biological processes at the turn of the 20th century. Odell was an equally gifted builder; upon meeting Foe, he set to work making her a microscope so she could film and track the oscillations of cell division in fruit fly embryos. Working together at the Center for Cell Dynamics at Friday Harbor, an interdisciplinary initiative that Odell would go on to direct, the two continued to collaborate closely. Among other projects, they constructed an experimentally validated mathematical model of how cytoskeletal fibres called microtubules are assembled to provide the scaffolding for a cell to divide.
Every time a human cell cycles, its genome can suffer 10 to 50 double-strand breaks – snapping the DNA helix clean through. When two breaks occur on the same chromosome, large stretches of DNA can be rearranged – or even fail to make it cleanly into a daughter cell – when the cell divides. Given this peril, our cells – and those of other multicellular species – have evolved exquisite mechanisms for DNA repair. Yet repair is not the same as restoration: these molecular toolkits can make mistakes, stitching together the wrong ends or reinserting fragments in the wrong place in the genome. Unlike point mutations, which alter DNA in small, local increments, double-strand breaks can rewrite the genome on a much larger scale. The nearly one-half of the human genome that is contained within genes – with some verging on the size of entire bacterial genomes – is at risk of break and misrepair.
Over the course of a lifetime, there are trillions of opportunities for the carefully timed rhythms in our bodies to break down. These vulnerabilities drive ageing and cancer by triggering cells to proliferate at the wrong time and place. ‘My husband’s death from cancer highlighted for me how it is that individuals die very often of damage to their genomes. They don’t die of something mechanical first,’ Foe tells me. But, she wonders, ‘why don’t we pass cancers on to offspring, or very rarely do?’ Given the fragility to which complexity has exposed our genes, Foe realised, there must be a way of preserving our bodies’ mind-bending intricacies. Somewhere between one generation and the next, there must be a checkpoint: a way to refuse broken instructions, to keep genetic inheritance intact enough to build again.
Organisms need a way of recognising one another to ensure they mate with the correct species
Sexual reproduction is one answer to that problem. Within all sexually reproducing species, a special kind of cell division called meiosis produces the cells – and the genetic material – that will be passed on to the next generation. During meiosis, each pair of chromosomes within the cell must first find each other and lock precisely together. In an elaborate dance of connection that can take several days in animals, and weeks of sloshing and jiggling in plants, the matching chromosomes zip themselves together until their DNA threads are aligned and bound together by a biological ‘glue’ called the synaptonemal complex.
If either chromosome is missing genes or has a chunk of DNA in the wrong place due to break or misrepair, the cell in which it resides is fated to die. Only chromosomes with aligned sequences can come apart again and partition into daughter cells that will become spores (fungi), seeds or pollen (plants), or eggs and sperm (animals).
This molecular embrace – called the pachytene checkpoint – has had an enormous impact on the evolution of life over the past 2.5 billion years. Crucially, Foe proposes, it keeps genetic timing mechanisms that maintain our cellular rhythms intact across generations. The pachytene checkpoint also explains why it is that closely related species, even when they can physiologically mate, produce offspring that are sterile; the DNA rearrangements that distinguish species, even when minor, trip the checkpoint during meiosis, preventing reproductive cells from being made. Species recognition is thus essential for survival. In dense ecosystems, such as the canopies of tropical rainforests where birds and insects fly en masse, organisms need a way of recognising one another to ensure they mate with the correct species and produce fertile offspring. Rather than a signal of fitness, Foe argues, the elaborate plumage of birds of paradise or the courtship dance of bowerbirds have evolved as wayfinding signals.
Like all things in life, the pachytene checkpoint is not 100 per cent effective. Things slip through, allowing organisms to adapt to changing environments and, Foe reasons, new species to emerge. Every once in a while, a break and misrepair will relocate a piece of DNA containing intact genes particularly relevant to the moment. By chance, this genetic shuffling escapes the ancient choreography of chromosomal aligning and checking during meiosis, and gets passed on to the next generation. This rearranged DNA tunes cellular rhythms, both posing a threat to fertility yet setting new boundaries for an organism’s capacities. ‘Say you’re a sand dune sunflower,’ Foe says, gesturing to the world outside the lab. In this flower, inherited misrepaired DNA has allowed its roots to become longer and its body more resistant to temperature fluctuations. It’s a hot summer and most sunflowers are making their way on the shady side of the dunes where the ground is consistently cooler. But the shuffled flower is thriving in the front. ‘Say there’s a couple of hot summers like this. Even though it doesn’t reproduce as well, it may gradually be able to establish itself.’ Having slipped through a protective mechanism, this flower gradually establishes a proto-species, one better adapted to its environment. The checkpoint then kicks back in to keep it, and its offspring, reproductively separate from parental species, carving a new evolutionary path.
Foe sees this latest work as an antidote to the long divide between molecular biology and whole-organism thinking, carrying us from the nucleus’s intimate mechanisms to evolution’s expansive sweep across the space and time of living forms. Only a broad view, she believes, can make sense of how complex life works. The lengthening of our genes, the enmeshing of temporality right into their fibres, made them vulnerable to potentially catastrophic damage. Dealing with this vulnerability has had magnificent side effects: ‘Here we are in the middle of this absolutely extraordinary planet and all of these creatures, and here’s a mechanism that just drives diversity. It drives beauty. We wouldn’t be here if it weren’t for that,’ Foe says. And as for biologists, ‘it’s been too long that you haven’t been talking to one another.’
The last time I spoke with Foe, she was preparing for the journey from San Juan to Denman Island, where she plans to move full time. The house is not yet finished but is liveable. It needs only, she tells me, a small utility room for firewood, tools, and a table for making botanic block prints inspired by the years her family spent in Mexico during her childhood. ‘Why block prints?’ I ask. To which Foe responds: ‘I like their aesthetics. I like the boldness of the region.’
For her next chapter, Foe wants to live simply: ‘I’ve got my garden planted except for beans and squash. I love having a garden again.’ For Foe, gardening is another way into the marvel that ‘you can have a seed, and it’s got all the information needed to make this huge, amazing plant.’ She wants to spend her remaining time planting and pruning as a way of ‘celebrating what it means to be a creature on this planet’. When I enquire what will become of her microscope in Friday Harbor, Foe seems ready to let it go. It’s not working properly, and she is unsure if she wants to repair it. ‘It could be a blessing in disguise,’ she admits reluctantly. ‘The truth of the matter is I just love looking down it and asking things, but I’m extremely happy on Denman Island.’
When not tending her garden, Foe’s priority will be getting the word out about genetic fragility and her hypothesis about the solution millions of years of evolution have wrought to protect the complexity of life. This project’s current form, an extended exposition in the journal Integrative Organismal Biology, is up against the reduced attention spans of modern readers. Foe’s prose resembles that of historical synthetic monographs such as Charles Darwin’s On the Origin of Species (1859) or D’Arcy Wentworth Thompson’s On Growth and Form (1917), which bore down on details over hundreds more pages. But in our fast-paced digital age, few scientists want to attend to long explanations any more, even when the intricacy of the things being described demand them. Foe is trying to break up this transformative story into digestible bites, but I can sense that it pains her to do so. Her other challenge is that the independent path she chose means a younger generation of biologists is largely unaware of her. Her focus has been on the work itself, cultivating life in and outside the lab, and not the scientific seminar or conference circuits.
The person Foe most wants to tell her story to, however, is Odell: ‘One of my greatest sorrows is that I never got to tell him what I think the answer is.’ As she envisions the ability to do so, she smiles and her whole being brightens: ‘He would’ve been delighted with it.’






