Where in the embryo does the person reside? Morphogenesis – the formation of the body from an embryo – once seemed so mystifying that scholars presumed the body must somehow already exist in tiny form at conception. In the 17th century, the Dutch microscopist Nicolaas Hartsoeker illustrated this ‘preformationist’ theory by drawing a foetal homunculus tucked into the head of a sperm.
This idea finds modern expression in the notion that the body plan is encoded in our DNA. But the more we come to understand how cells produce shape and form, the more inadequate the idea of a genomic blueprint looks, too. What cells follow is not a blueprint; if they can be considered programmed at all, it’s not with a plan of what to make, but with a set of rules to guide construction. One implication is that humans and other complex organisms are not the unique result of cells’ behaviour, but only one of many possible outcomes.
This view of the cell as a contingent, constructional entity challenges our traditional idea of what a body is, and what it can be. It also opens up some remarkable and even disconcerting possibilities about the prospects of redirecting biology into new shapes and structures. Life suddenly seems more plastic and amenable to being reconfigured by design. Understanding the contingency and malleability of multicellular form also connects us to our deep evolutionary past, when single-celled organisms first discovered the potential benefits of becoming multicellular. ‘The cell may be the focus of evolution, more than genes or even than the organism,’ says Iñaki Ruiz-Trillo of the Institute of Evolutionary Biology in Barcelona. Far from the pinnacle of the tree of life, humans become just one of the many things our cells are capable of doing.
In one of the most dramatic demonstrations to date that cells are capable of more than we had imagined, the biologist Michael Levin of Tufts University in Medford, Massachusetts and his colleagues have shown that frog cells liberated from their normal developmental path can organise themselves in distinctly un-froglike ways. The researchers separated cells from frog embryos that were developing into skin cells, and simply watched what the free cells did.
Culturing cells – growing them in a dish where they are fed the nutrients they need – is a mature technology. In general, such cells will form an expanding colony as they divide. But the frog skin cells had other plans. They clustered into roughly spherical clumps of up to several thousand cells each, and the surface cells developed little hairlike protrusions called cilia (also present on normal frog skin). The cilia waved in coordinated fashion to propel the clusters through the solution, much like rowing oars. These cell clumps behaved like tiny organisms in their own right, surviving for a week or more – sometimes several months – if supplied with food. The researchers called them xenobots, derived from Xenopus laevis, the Latin name of the African clawed frog from which the cells were taken.
A swarm of living robots (xenobots) swimming through water, time lapse. Courtesy Douglas Blackiston
Some of this wasn’t entirely new. Scientists have known for more than a century that a piece of embryonic tissue destined to become skin will, if cut off and cultured, grow cilia. Such a piece of tissue is called the ‘animal cap’, and various studies have shown that Xenopus animal caps can, if given the right biochemical signals, grow into many other tissue types, including neurons, muscle and even beating heart tissue.
But Levin and colleagues now say that some of these structures are not just random blobs of sticky cells: they resemble autonomous organisms. If they are damaged, the cells heal back in the original shape. They can signal to one another by emitting pulses of calcium ions, although the researchers aren’t sure what the message conveys. They move with apparent purpose, sometimes circling one another or sweeping up other individual cells around them into piles.
The cells are like Lego bricks that can be assembled in different ways – except they do the assembly themselves
These xenobots seem to represent an entirely different developmental program, Levin says, that the frog cells can adopt. Having been freed from their usual environment, it’s as if the cells are able to discover a new way of life. What’s baffling is that they are genetically no different to ordinary frog cells. So what does the genome encode, if not a ‘plan for a frog’?
It seems that, instead, the genes are part of a molecular program that gives cells certain tendencies, for example to stick together in particular configurations. The cells are like Lego bricks that can be assembled in different ways – except that the cells do the assembly themselves. In the environment experienced by a normal embryo, the principles for assembly generate first a tadpole and then a frog. But these aren’t the only possible solutions to the collective computation that the cells perform. Xenobots are another one, and perhaps there are more morphologies – body structures – yet to be discovered.
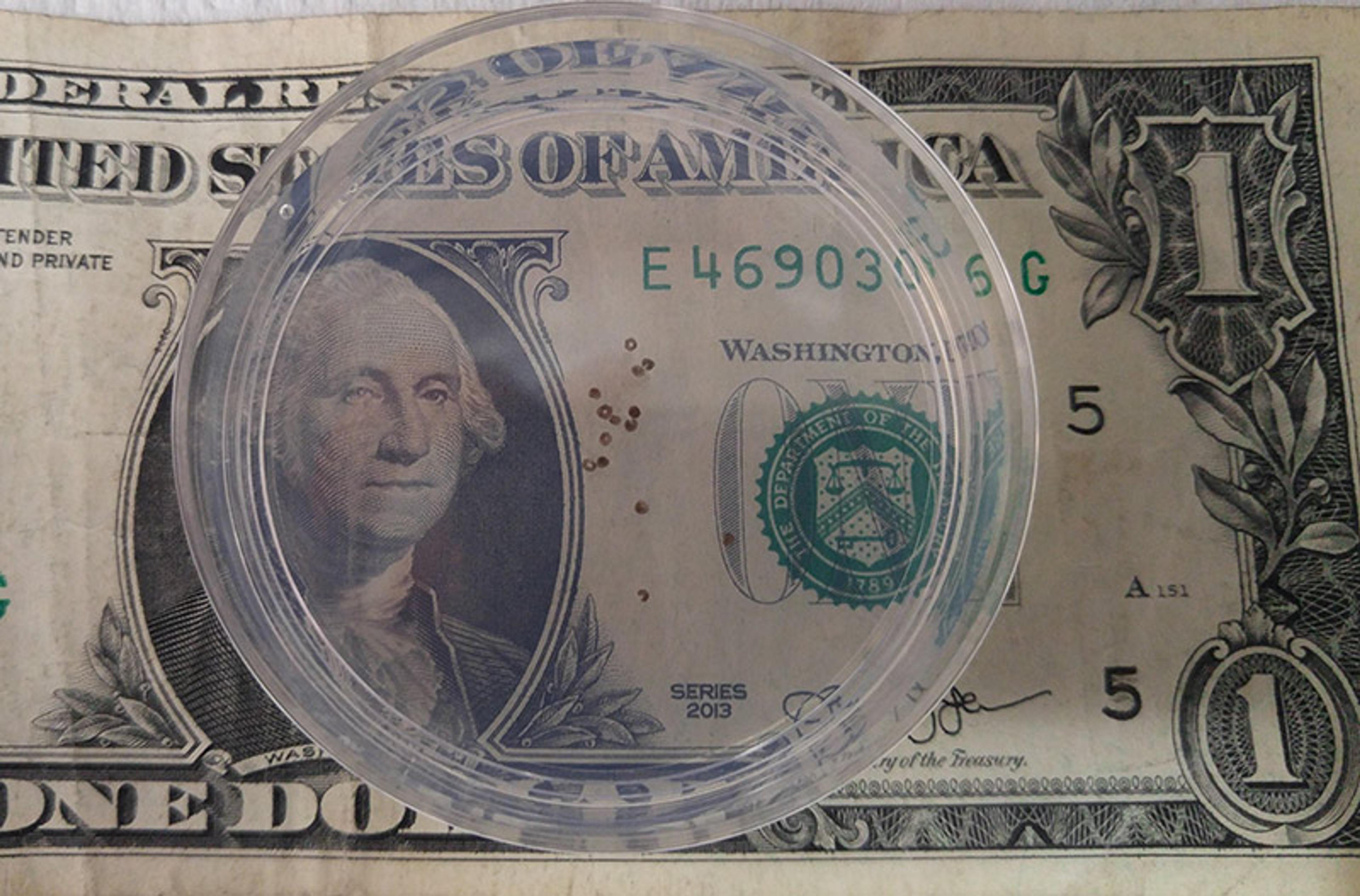

A group of AI-designed organisms in a Petri dish (dollar bill for scale). Courtesy Douglas Blackiston
Levin and colleagues have recently found a new type of behaviour that xenobots can exhibit. They discovered that these pseudo-organisms can even replicate, after a fashion. Xenobots placed in a dish of cells will move to marshal those loose cells into piles that, over the course of a few days, cluster into new xenobots that then take off through the liquid themselves. Left to their own devices, the xenobots typically manage to produce only a single generation of offspring. But the researchers wondered if they could do better. They made computer simulations to search for xenobot shapes that were better at making new xenobots, using an AI program devised by their team member Josh Bongard of the University of Vermont. The simulations suggested that structures like C-shaped half-doughnuts could sweep up cells more efficiently than the spheroidal xenobots could, making larger (spherical) clusters of ‘offspring’.
The team then shaped such xenobots by hand, using tiny tools to manipulate the cells into the C-shaped forms, and then letting them go to work on dishes of fresh cells. Because the resulting offspring were bigger than those of earlier xenobots, they were able to sustain the replication process over more generations. Whereas the best spheroidal parent xenobots could ever do is make two generations before the offspring become too small to reproduce further, the C-bots could engender four further generations of offspring, each of them spheroidal and gradually decreasing in average size.
This is not, of course, how real living organisms reproduce. There’s no cell division involved for xenobots, so the parents don’t pass on genetic material to the offspring. Rather, the behaviour looks more like that seen in some robots made from reconfigurable parts, which have been shown to be capable of assembling copies of themselves when supplied with the parts. The xenobots can’t evolve this way, and have to be given the parts. Levin and colleagues call the process kinematic self-replication: making copies by movement rather than by biological reproduction. ‘Replication is what’s available to entities that cannot absorb new mass themselves,’ says Bongard. ‘They can behave, but not grow. Such systems can only combine objects in their external environment into copies of themselves.’
The work shows that, by combining biological xenobots with the exploratory power of AI, it’s possible to make a kind of ‘living machine’ devised for a purpose. ‘AI can be brought in to exaggerate an innate capability,’ says Bongard. ‘The AI can “program” new behaviours into organisms by rearranging their morphology rather than their genes.’ The researchers wonder if the simulations might identify other shapes that can assemble different structures, or perhaps perform other tasks entirely. ‘One of my primary interests in this project is exactly how ‘far’ from the wild type [the natural, spontaneously arising form of xenobots] an AI can push things,’ says Bongard. ‘We’re now working on incorporating several new behaviours into xenobots via AI-driven design.’
This perspective entails a new way of thinking about cells: not as building blocks assembled according to a blueprint, but as autonomous entities with skills that can be leveraged to make all manner of organisms and living structures. You might conceive of them as smart, reprogrammable, shapeshifting robots that can move, stick together, and signal to one another – and, by those means, build themselves into elaborate artefacts.
This might also be a better way to conceptualise how our own bodies are built during embryogenesis. That growth process is a gradual and sequential unfolding of shape in which each step creates the conditions for the next. Crucially, this unfolding involves changes in the state of the cells themselves, triggered by signals from their surroundings. For example, as a sheet of cells grows in a tissue that is confined at its edges by the rest of the embryo, the sheet is forced to buckle. The mechanical forces at the tip of a bulge might then be detected by sensors on the cell surfaces, which in turn triggers a chemical signal that’s relayed to the genes. Genes are then switched on so as to alter the cells’ properties – making them less sticky and more mobile, say, creating new options for the shape of the tissue. There is a continual dialogue between what is happening outside the cell and what is going on inside, between the shape and form of the overall structure of the growing embryo and the genetic activity inside its components.
This rich and subtle conversation can make it hard to predict which shapes cells such as ours will grow into – something rendered more difficult by the fact that, unlike those of bacteria, our cells alter their gene activity permanently as they divide and grow from general-purpose embryonic stem cells into specialised tissues, switching some genes off and turning others on. A new discipline called synthetic morphology seeks to accommodate and even exploit this complication in order to harness the constructive capacities of cells to make entirely new, non-natural, multicellular structures and devices. If it’s going to be capable of predicting and directing the outcomes, the endeavour will most probably rely on the kind of AI and other computational resources deployed by Levin and colleagues.
The most noticeable aspects of organisms – their overall shape and form – might be the most ‘shallowly’ encoded
It’s not too far-fetched to suppose that synthetic morphology includes the possibility of making entirely new organisms. Arguably xenobots are already such a creature, although these microscopic blobs are not much to look at. Could possible solutions to the assembly computation of frog cells – what mathematical biologists call attractor states – include totally different macroscopic bodies: fish-like, say, or worm-like?
‘We are asking AI to tell us more about the attractor landscape of X laevis,’ says Bongard. ‘The AI has found two such attractors so far. How many other attractors are out there to be discovered? And how many exist for other species?’
That thought might defy our intuitions about how life works, but the truth is that the most noticeable aspects of organisms – their overall shape and form – might in fact be the most ‘shallowly’ encoded, determined not so much by deep genetic resources as by the way the assembly rules happen to play out in any given instance. In some sense, the very existence of tadpoles and frogs (let alone xenobots) as viable outputs of the Xenopus genome testifies to that hypothesis: tadpoles are not morphologically mini-frogs in the way that babies are mini-adults, but they ‘work’ as organisms in their own right. They’re an attractor in which frog cells can temporarily reside.
The revelations about cells’ morphological potential shows why the fact we’re so genetically similar to our evolutionary relatives – having 99 per cent genetic overlap with chimpanzees and 84 per cent with dogs – kind of misses the point. Most of our genetic resources seem to be there to create and sustain the constructive capabilities of the cells that make us. Exactly which forms they build can look like an afterthought: from this perspective, all it requires to make the difference between a human body and a dog body is a little fine-tuning of the rules that govern development. Of course, those differences are vital to evolutionary success in the niche that an organism inhabits but, even so, morphology is lightly written into the developmental script.
This is indeed what evolutionary genetics seems to be telling us. There hasn’t been much innovation in genes themselves since the earliest complex multicellular organisms – the metazoans – appeared around 750 million years ago. Our shock at discovering, thanks to the Human Genome Project, that we have only about as many protein-coding genes as the tiny soil-dwelling worm Caenorhabditis elegans (which has just 2,000-3,000 cells in its body) might have been largely due to a sense of hurt pride that we are not so special after all. But perhaps it was an indication that we had the wrong view of what those genetic resources were for in the first place: they don’t supply the plan, but help to create options.
After all, most of the genes and capabilities needed for a multicellular lifestyle were present even in our single-celled ancestors. They already possessed the ability to send signals to one another to enable cooperative behaviour, to stick together, and to specialise into different cell types. We can see such abilities today in single-celled amoebae such as the slime mould Dictyostelium discoideum, which can assemble into multicellular ‘bodies’ under stress.
Ruiz-Trillo and his coworkers think that the genetic resources required for such multicellular behaviour stem largely from mechanisms for regulating genes – turning them on and off – rather than from any novelty in the genes themselves. ‘Much of the innovation in gene content seen in the transition to multicellularity is rooted in pervasive “tinkering” with pre-existing gene families,’ they say. The researchers were led to this conclusion from their studies of the amoeba Capsaspora owczarzaki, one of the closest evolutionary relatives of the first multicellular animals, which will have resembled sponges and comb jellies. Capsaspora has more genes involved in regulatory functions than any other single-celled organism, mostly encoding proteins called transcription factors. Ruiz-Trillo and colleagues discovered that the biomolecular interaction networks that these proteins govern in Caspaspora are often found in animals too. In other words, the networks were already ‘primed and ready to go’ before true multicellularity took off.
In some ways, tumours represent an alternative morphology of our own cells
In one sense, our continued proximity to the boundary of unicellular and multicellular lifestyles might be seen as the reason we (and nearly all metazoans) are prone to cancer, in which our cells seem to have abandoned the restraint required for multicelled living and to have returned to the proliferating excess of the single cell. ‘Probably the most important problem of being multicellular is the “cheating” of some cells that decide to go by themselves,’ says Ruiz-Trillo. ‘A lot of the genes that are crucial for animal multicellularity are genes involved in cancer disease. Perhaps being multicellular may be against our ancestral mode of living, and something that takes continuous effort and energy to maintain.’
The flipside of that view is that even single-celled living is apt to become collective. Not even cancer cells are bent on self-replication, oblivious to other cells around them. Many tumours look less like undifferentiated masses of wildly multiplying cells and more like a deranged version of organ growth. Cancer cells, too, can differentiate and specialise, as if following some new, lunatic trajectory. A tumour, far from growing heedless of the host tissues around it, can integrate with those tissues and even commandeer them for its own ends. In some ways, tumours represent an alternative morphology of our own cells.
This same pattern of evolutionary innovation through novelty in gene use, rather than in genes themselves, continued as multicellular life got more complex. The biologist Michel Morange of the École Normale Supérieure in Paris says that ‘the major changes observed during evolution are more the consequence of the reorganisation of [gene regulatory] networks than the modification of the protein links that form them.’ In 2011, the developmental biologists Craig Lowe, David Haussler and their colleagues investigated what kinds of regulatory changes have been involved in the evolution of vertebrates since their first appearance about 650 million years ago. They compared the genomes of various vertebrates – humans, cows, mice, and two types of fish (sticklebacks and medaka, or Japanese rice fish) – to see which genetic sequences they shared, and which their common ancestors presumably had too.
The researchers considered parts of the sequences not usually examined in such phylogenetic comparisons: so-called nonexonic elements, which fall outside of the sequences that encode proteins. Nonexonic sequences are often assumed to be random genomic junk accumulated by accident, but Lowe and colleagues reasoned that if some nonexonic elements were found to be highly conserved – to recur more or less unchanged in different species – then they probably have some functional role in the cell. This means they would be subject to selection pressure, which will preserve them, whereas random junk would be expected to degenerate quickly and diverge in sequence between different species. The researchers figured that such conserved nonexonic elements (CNEEs) will probably be involved in regulating the activity of genes.
In the Cambrian explosion around 540 million years ago, all manner of strange body shapes appeared
They found that, rather than smooth and gradual changes in the frequencies of CNEEs, three distinct eras of change seem to have occurred since vertebrates first evolved. Until about 300 million years ago, when mammals split from birds and reptiles, changes in regulation seem to have happened mostly in parts of the genome close to transcription factors and the key genes that they control. Then, between 300 and 100 million years ago, those changes tailed off; instead, modifications were observed near genes that code for the protein molecules serving as signal receptors at the cell surface. In other words, what seemed to matter for these evolutionary changes was a shift not in the content of cells but the way they talk to one another: the dialogue that makes multicellularity possible. Finally, as of 100 million years ago, during the emergence of placental mammals (that is, all mammals except marsupials and monotremes such as echidnas), the regulatory changes seem to be associated with mechanisms for modifying the structure of proteins after they have been synthesised in raw form, especially for proteins that are associated with transmitting signals within cells.
Evolution, then, might be considered to have successively discovered ways to innovate and generate new organisms by reshuffling, first, how developmental genes are switched on and off, then how cells communicate, and finally how information gets passed around inside cells. In all cases, the action is focused on how cells interact with and respond to one another, rather than what individual cells do: in other words, they are changes to the multicellular assembly rules. ‘It appears that our set of around 20,000 genes is capable of building morphologies that wildly vary in both their shape and complexity,’ says Lowe. ‘We think that complexity is encoded by how many rules are encoded in the genome to control when and where those genes turn on and off.’ The C elegans worm, while having roughly as many genes, has rather little of this regulatory sophistication, and so is stuck in a low-complexity form.
The generative potential of cells equipped for multicellular construction was evident almost as soon as this became a lifestyle option, in evolutionary terms. In the Cambrian explosion around 540 million years ago, all manner of strange body shapes appeared, many of which are no longer exhibited by any creatures on Earth. Perhaps we should regard those forgotten ‘endless forms most beautiful’, to borrow Charles Darwin’s resonant phrase, as an illustration of the constructive potential of the metazoan cell – an exuberant expression of the palette of solutions to the problem of cell assembly, which natural selection then stringently pruned.
Acknowledging that the human form is a contingent outcome of the way our cells are programmed for construction raises some mind-bending questions. Are there, for example, human xenobots (perhaps we might call them anthrobots)? If so, are they truly ‘human’? Might there be a kind of organ or tissue that our cells could make but don’t normally get the chance to? Might our still cells ‘remember’ older evolutionary body shapes?
Perhaps at least some of the attractors that AI algorithms could help identify in xenobot-type assemblies ‘are echoes of past targets of natural selection’, says Bongard. ‘At least some of these new configurations may tell us things about the environments and selection pressures that acted on these organisms in their distant past, and how they responded evolutionarily. In a way, these attractors are like fossils: they might be giving us partial glimpses into the past.’
They also raise questions about how much we can reshape biological forms – including our own. In one sense, we already know that the human body has considerable ‘plasticity’. No genetic signal tells an embryo to split into identical twins, for example: it’s just one way the assembly rules happen to play out. And even relatively modest genetic tweaks to the rules can generate markedly different bodies. The heritable condition called Kartagener’s syndrome, for example, which leads to respiratory problems in early childhood, can sometimes be accompanied by complete mirror-image reversal of the internal organs – the heart lying to the right, say – which leads to health complications. It’s as if one key step in the early formation of the body plan went awry but then the cells accommodated it as best they could. Developmental problems such as spina bifida, in which the neural tube that will become the spinal cord fails to close, have many complex and imperfectly understood causes that may all lead to the ‘wrong’ outcome of the assembly rules.
‘Wrong’, that is, for the health of the baby. But sometimes the rules result in a morphological outcome only a little different from the most common one: an extra finger, a shorter limb or overall stature, a cleft palate. These are only ‘errors’ if we choose to make them so; sometimes, a non-standard body shape is only a problem because, as a society, we don’t make adequate allowance for it, physically, aesthetically or socially. There are many ways that humans can look, because the result isn’t written into our genome. What cells get are guidelines for assembly, and the result is not prescribed. Nature has realised that there are better and more versatile ways to build than that.